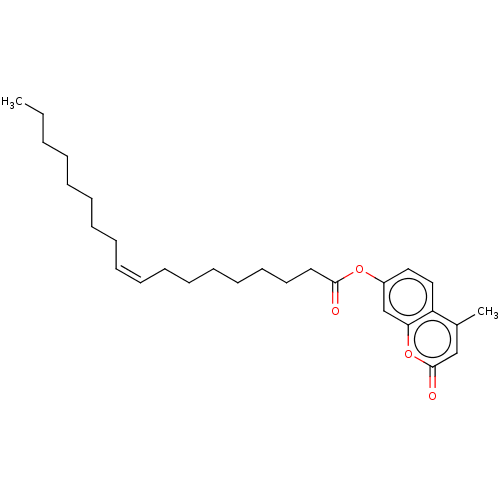
 BDBM246504 4-Methylumbelliferyl oleate (4-MU oleate)
BDBM246504 4-Methylumbelliferyl oleate (4-MU oleate)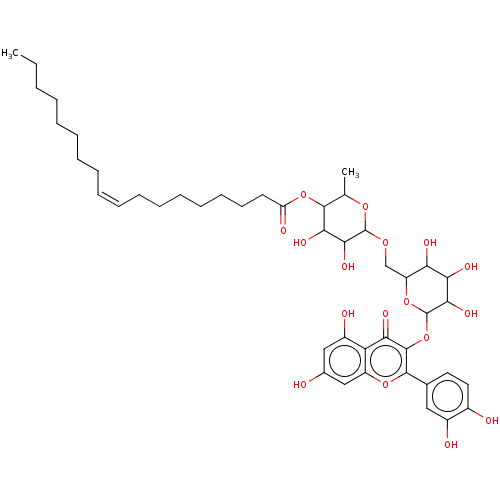 BDBM237193 Rutin oleate
BDBM237193 Rutin oleate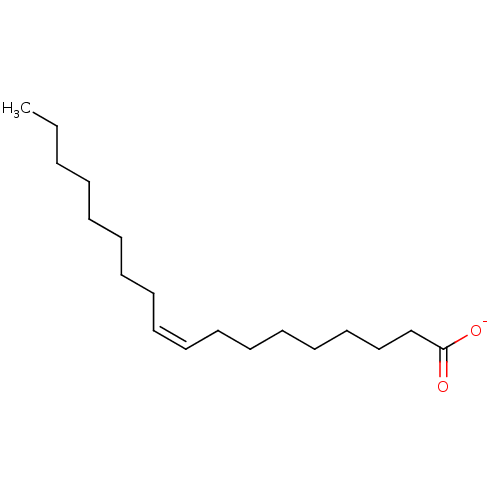 BDBM50448437 CHEMBL3122151 Oleate
BDBM50448437 CHEMBL3122151 Oleate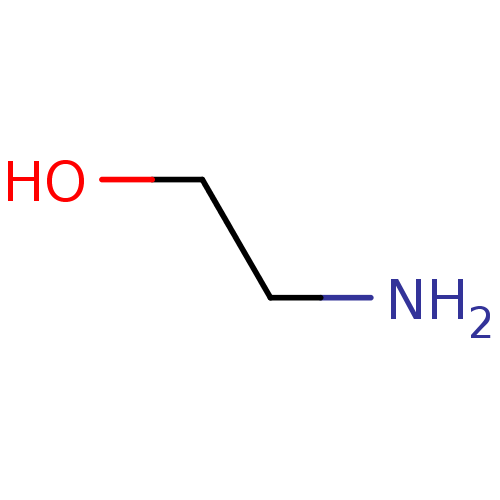 ethanolamine BDBM7973 CHEMBL104943 2-aminoethan-1-ol
ethanolamine BDBM7973 CHEMBL104943 2-aminoethan-1-ol 2-(benzylamino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 1 BDBM18072
2-(benzylamino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 1 BDBM18072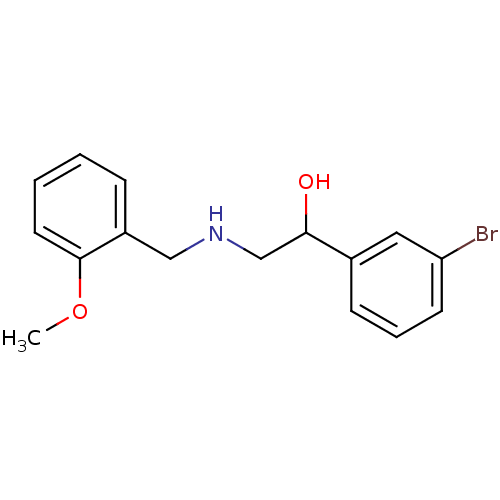 1-(3-bromophenyl)-2-{[(2-methoxyphenyl)methyl]amino}ethan-1-ol BDBM18082 ethanolamine, 18
1-(3-bromophenyl)-2-{[(2-methoxyphenyl)methyl]amino}ethan-1-ol BDBM18082 ethanolamine, 18 2-{[(2-methoxyphenyl)methyl]amino}-1-(3-methylphenyl)ethan-1-ol BDBM18083 ethanolamine, 19
2-{[(2-methoxyphenyl)methyl]amino}-1-(3-methylphenyl)ethan-1-ol BDBM18083 ethanolamine, 19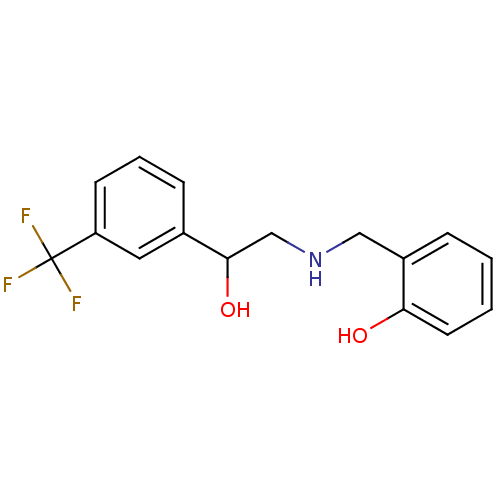 BDBM18077 ethanolamine, 9 2-[({2-hydroxy-2-[3-(trifluoromethyl)phenyl]ethyl}amino)methyl]phenol
BDBM18077 ethanolamine, 9 2-[({2-hydroxy-2-[3-(trifluoromethyl)phenyl]ethyl}amino)methyl]phenol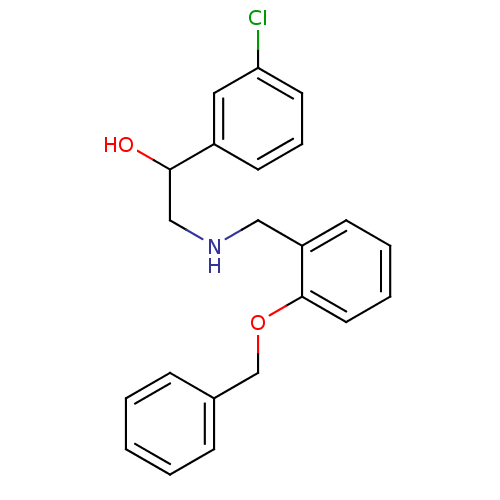 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-chlorophenyl)ethan-1-ol ethanolamine, 22 BDBM18085
2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-chlorophenyl)ethan-1-ol ethanolamine, 22 BDBM18085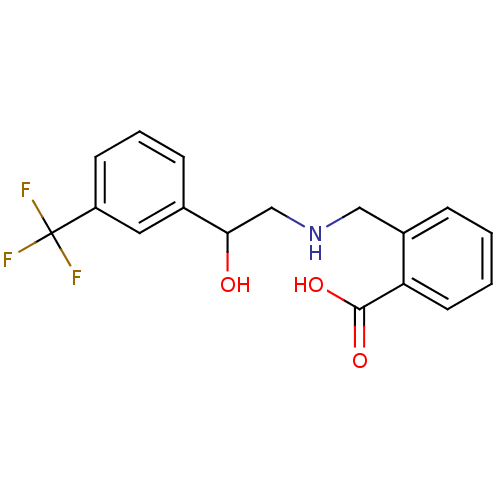 2-[({2-hydroxy-2-[3-(trifluoromethyl)phenyl]ethyl}amino)methyl]benzoic acid BDBM18075 ethanolamine, 7
2-[({2-hydroxy-2-[3-(trifluoromethyl)phenyl]ethyl}amino)methyl]benzoic acid BDBM18075 ethanolamine, 7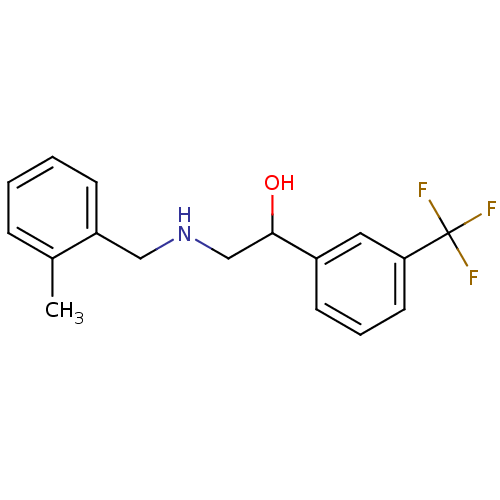 BDBM18074 ethanolamine, 6 2-{[(2-methylphenyl)methyl]amino}-1-[3-(trifluoromethyl)phenyl]ethan-1-ol
BDBM18074 ethanolamine, 6 2-{[(2-methylphenyl)methyl]amino}-1-[3-(trifluoromethyl)phenyl]ethan-1-ol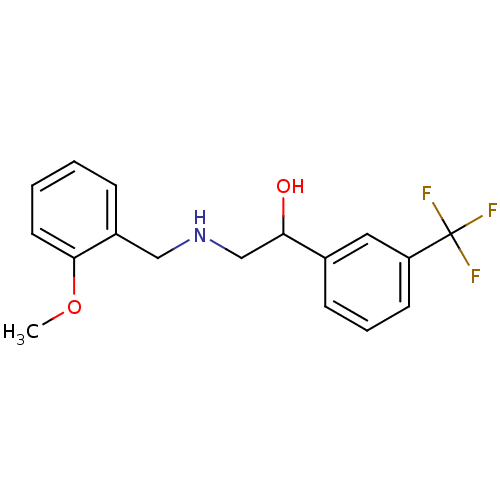 ethanolamine, 11 2-{[(2-methoxyphenyl)methyl]amino}-1-[3-(trifluoromethyl)phenyl]ethan-1-ol BDBM18079
ethanolamine, 11 2-{[(2-methoxyphenyl)methyl]amino}-1-[3-(trifluoromethyl)phenyl]ethan-1-ol BDBM18079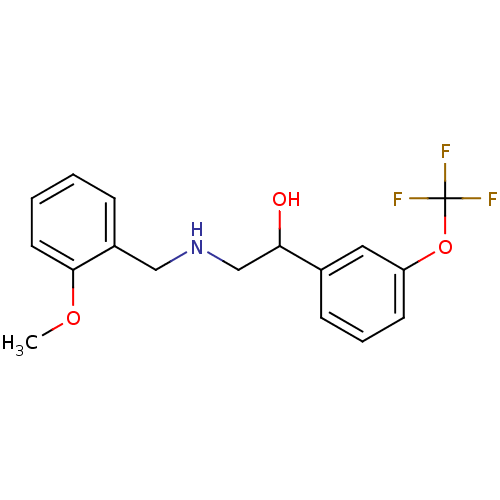 ethanolamine, 20 BDBM18084 2-{[(2-methoxyphenyl)methyl]amino}-1-[3-(trifluoromethoxy)phenyl]ethan-1-ol
ethanolamine, 20 BDBM18084 2-{[(2-methoxyphenyl)methyl]amino}-1-[3-(trifluoromethoxy)phenyl]ethan-1-ol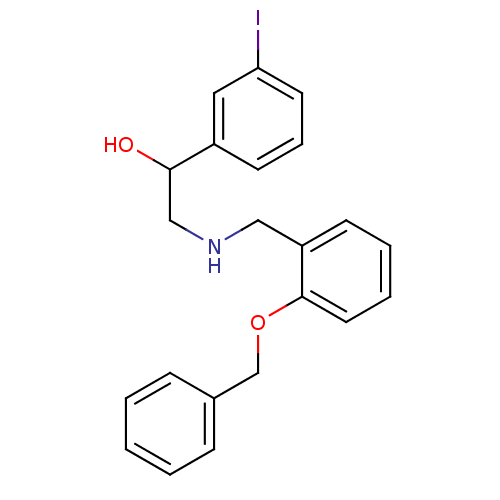 ethanolamine, 23 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-iodophenyl)ethan-1-ol BDBM18086
ethanolamine, 23 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-iodophenyl)ethan-1-ol BDBM18086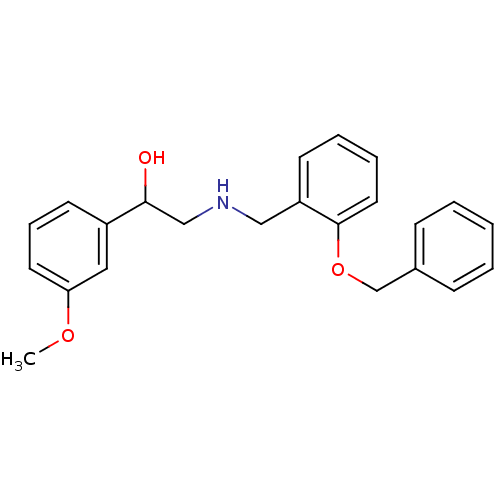 ethanolamine, 24 BDBM18087 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-methoxyphenyl)ethan-1-ol
ethanolamine, 24 BDBM18087 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-methoxyphenyl)ethan-1-ol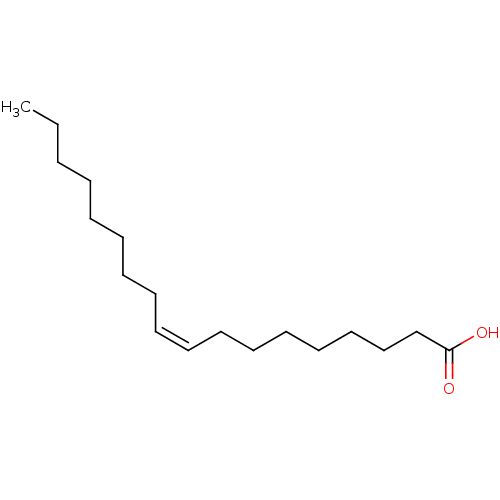 oleicacid (Z)-9-octadecenoic acid OLEATE BDBM50150484 Octadec-9-enoic acid (Z)-Octadec-9-enoic acid CHEMBL8659 oleaic acid
oleicacid (Z)-9-octadecenoic acid OLEATE BDBM50150484 Octadec-9-enoic acid (Z)-Octadec-9-enoic acid CHEMBL8659 oleaic acid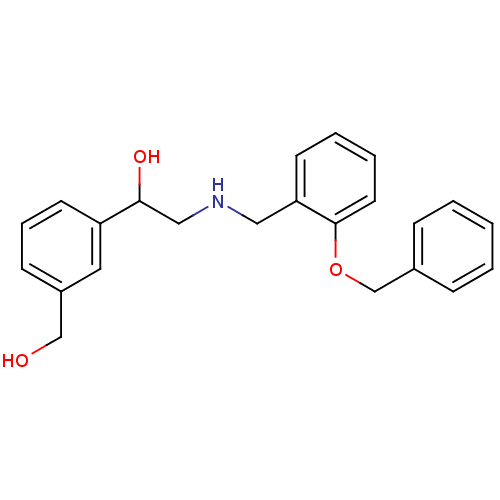 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-[3-(hydroxymethyl)phenyl]ethan-1-ol BDBM18088 ethanolamine, 25
2-({[2-(benzyloxy)phenyl]methyl}amino)-1-[3-(hydroxymethyl)phenyl]ethan-1-ol BDBM18088 ethanolamine, 25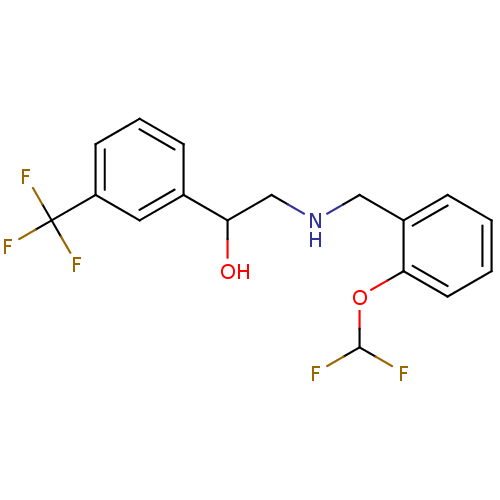 2-({[2-(difluoromethoxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 10 BDBM18078
2-({[2-(difluoromethoxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 10 BDBM18078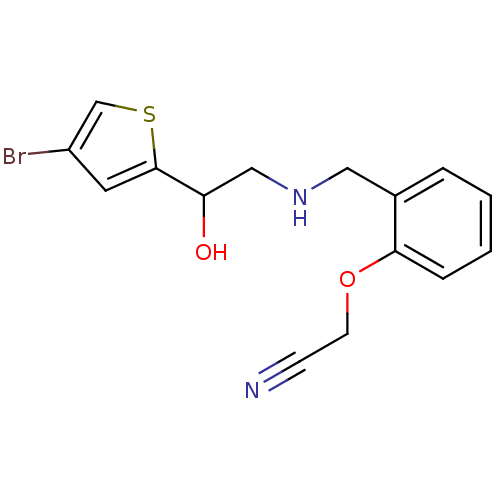 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]acetonitrile BDBM18092 ethanolamine, 37
2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]acetonitrile BDBM18092 ethanolamine, 37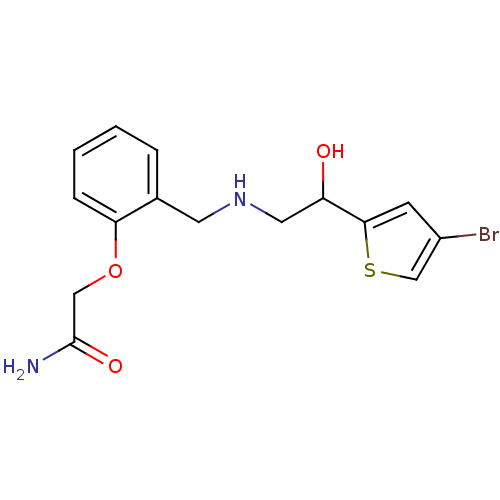 BDBM18104 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]acetamide ethanolamine, 49
BDBM18104 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]acetamide ethanolamine, 49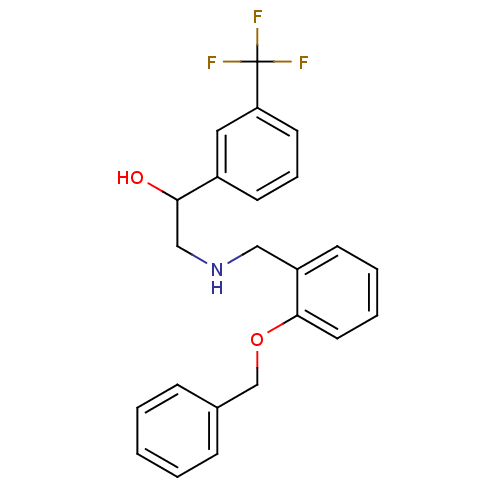 ethanolamine, 13 BDBM18081 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol
ethanolamine, 13 BDBM18081 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol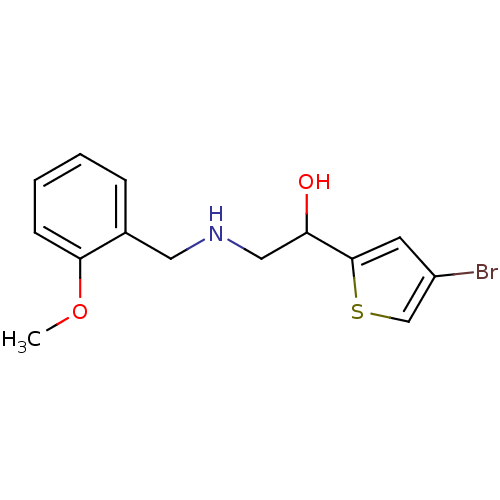 ethanolamine, 34 1-(4-bromothiophen-2-yl)-2-{[(2-methoxyphenyl)methyl]amino}ethan-1-ol BDBM18089
ethanolamine, 34 1-(4-bromothiophen-2-yl)-2-{[(2-methoxyphenyl)methyl]amino}ethan-1-ol BDBM18089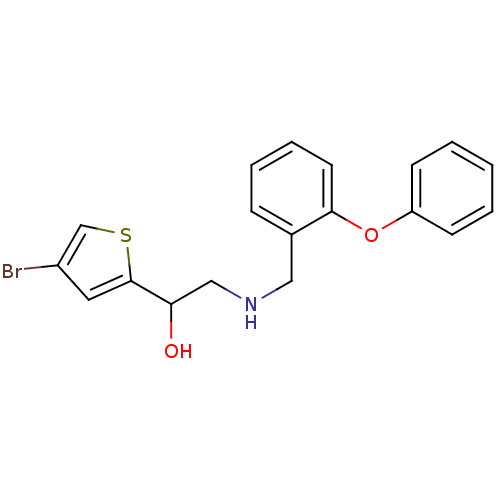 ethanolamine, 35 BDBM18090 1-(4-bromothiophen-2-yl)-2-{[(2-phenoxyphenyl)methyl]amino}ethan-1-ol
ethanolamine, 35 BDBM18090 1-(4-bromothiophen-2-yl)-2-{[(2-phenoxyphenyl)methyl]amino}ethan-1-ol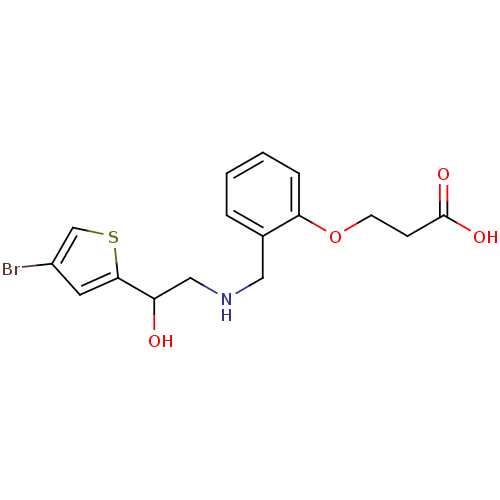 3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]propanoic acid BDBM18108 ethanolamine, 53
3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]propanoic acid BDBM18108 ethanolamine, 53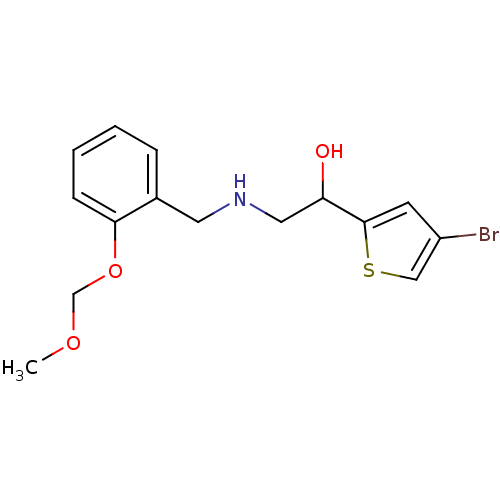 BDBM18091 1-(4-bromothiophen-2-yl)-2-({[2-(methoxymethoxy)phenyl]methyl}amino)ethan-1-ol ethanolamine, 36
BDBM18091 1-(4-bromothiophen-2-yl)-2-({[2-(methoxymethoxy)phenyl]methyl}amino)ethan-1-ol ethanolamine, 36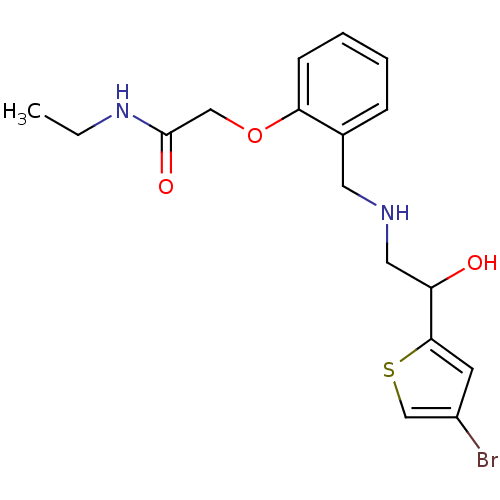 BDBM18105 ethanolamine, 50 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-ethylacetamide
BDBM18105 ethanolamine, 50 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-ethylacetamide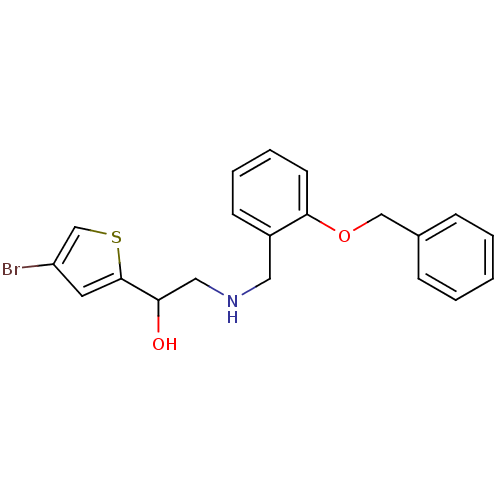 ethanolamine, 43 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol BDBM18098
ethanolamine, 43 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol BDBM18098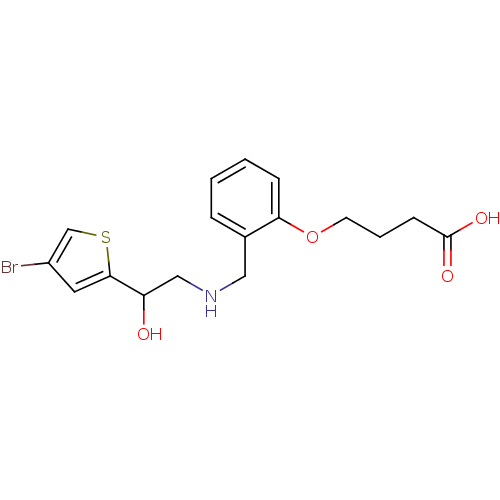 ethanolamine, 56 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]butanoic acid BDBM18111
ethanolamine, 56 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]butanoic acid BDBM18111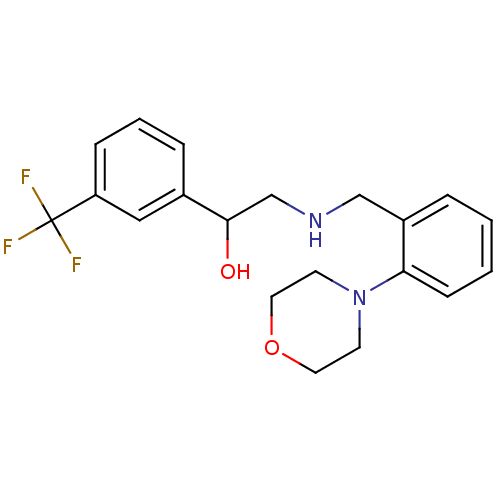 2-({[2-(morpholin-4-yl)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol BDBM18076 ethanolamine, 8
2-({[2-(morpholin-4-yl)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol BDBM18076 ethanolamine, 8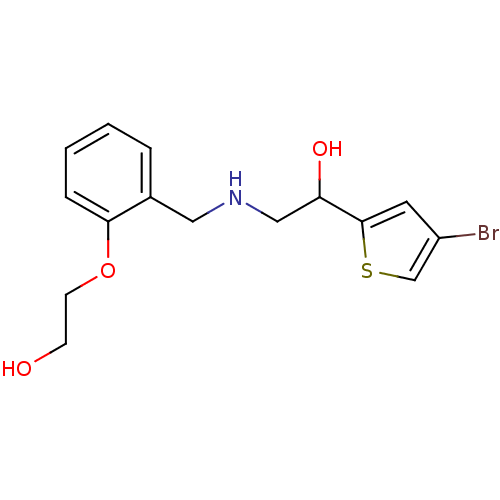 BDBM18093 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol ethanolamine, 38
BDBM18093 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol ethanolamine, 38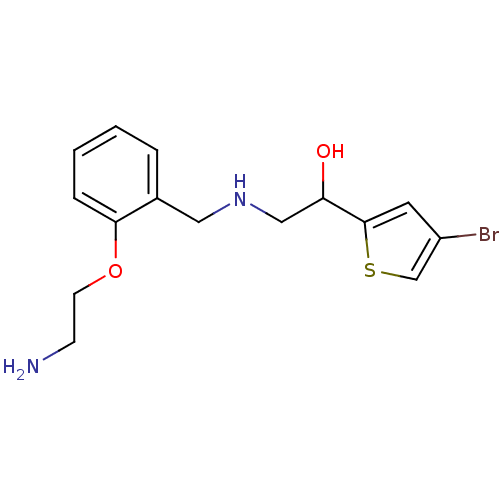 BDBM18094 ethanolamine, 39 2-({[2-(2-aminoethoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol
BDBM18094 ethanolamine, 39 2-({[2-(2-aminoethoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol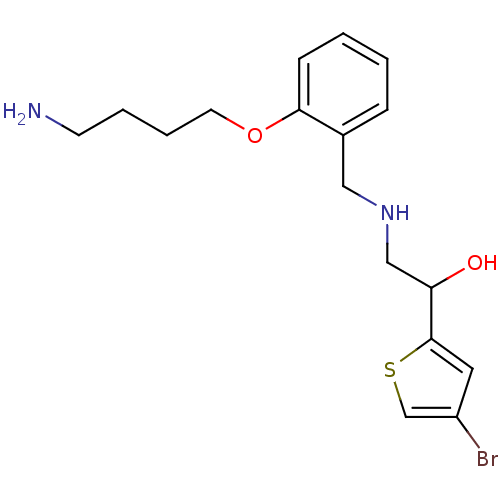 BDBM18114 2-({[2-(4-aminobutoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol ethanolamine, 59
BDBM18114 2-({[2-(4-aminobutoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol ethanolamine, 59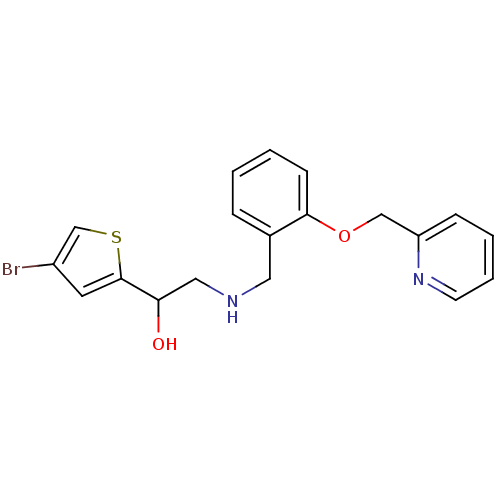 1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-2-ylmethoxy)phenyl]methyl}amino)ethan-1-ol ethanolamine, 45 BDBM18100
1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-2-ylmethoxy)phenyl]methyl}amino)ethan-1-ol ethanolamine, 45 BDBM18100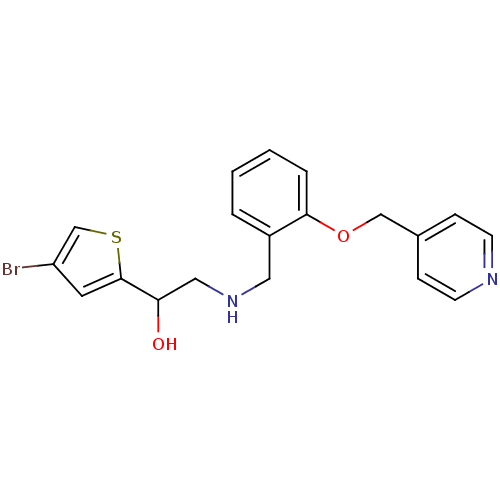 1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-4-ylmethoxy)phenyl]methyl}amino)ethan-1-ol BDBM18102 ethanolamine, 47
1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-4-ylmethoxy)phenyl]methyl}amino)ethan-1-ol BDBM18102 ethanolamine, 47 BDBM18106 ethanolamine, 51 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]acetamide
BDBM18106 ethanolamine, 51 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]acetamide BDBM18116 ethanolamine, 64 2-[2-({[(2R)-2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol
BDBM18116 ethanolamine, 64 2-[2-({[(2R)-2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol BDBM18117 ethanolamine, 65 2-[2-({[(2S)-2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol
BDBM18117 ethanolamine, 65 2-[2-({[(2S)-2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol ethanolamine, 44 BDBM18099 1-(4-bromothiophen-2-yl)-2-[({2-[(4-methanesulfonylphenyl)methoxy]phenyl}methyl)amino]ethan-1-ol
ethanolamine, 44 BDBM18099 1-(4-bromothiophen-2-yl)-2-[({2-[(4-methanesulfonylphenyl)methoxy]phenyl}methyl)amino]ethan-1-ol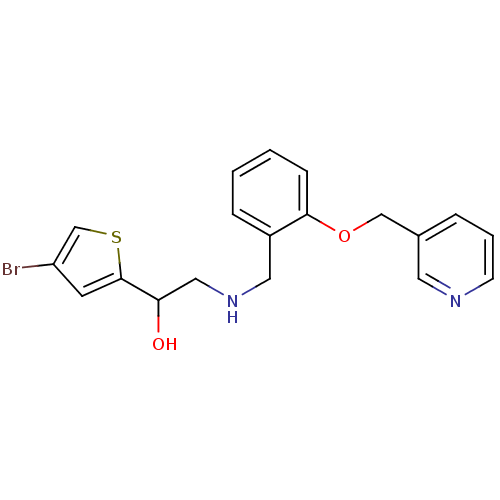 ethanolamine, 46 1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-3-ylmethoxy)phenyl]methyl}amino)ethan-1-ol BDBM18101
ethanolamine, 46 1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-3-ylmethoxy)phenyl]methyl}amino)ethan-1-ol BDBM18101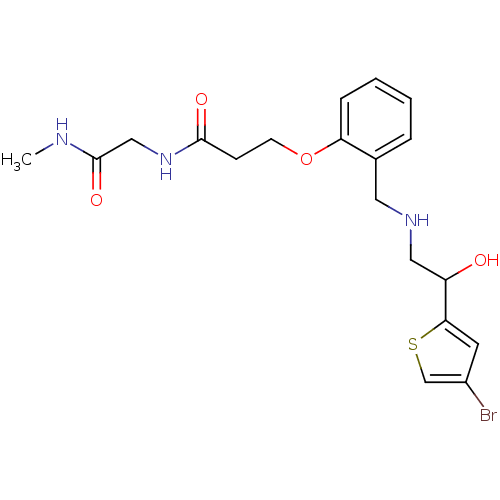 ethanolamine, 54 3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]propanamide BDBM18109
ethanolamine, 54 3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]propanamide BDBM18109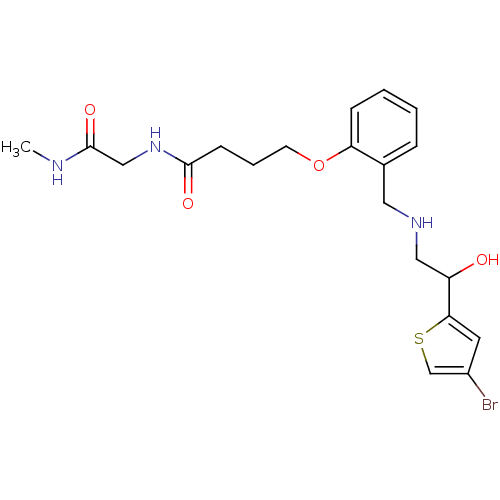 ethanolamine, 57 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]butanamide BDBM18112
ethanolamine, 57 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]butanamide BDBM18112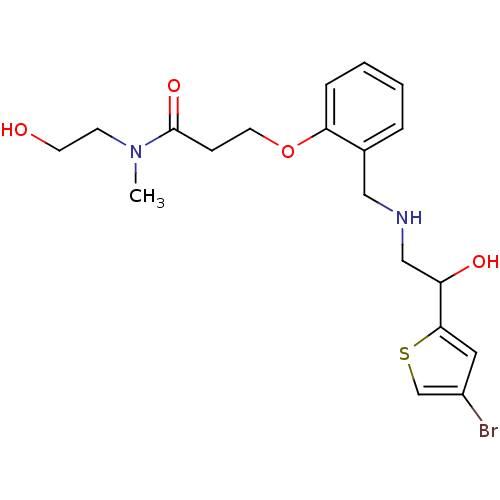 3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylpropanamide ethanolamine, 55 BDBM18110
3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylpropanamide ethanolamine, 55 BDBM18110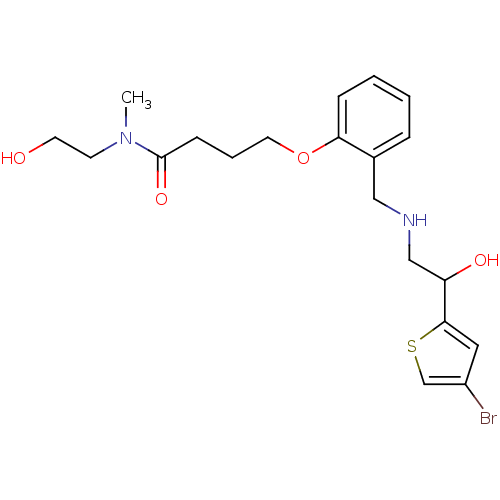 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylbutanamide BDBM18113 ethanolamine, 58
4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylbutanamide BDBM18113 ethanolamine, 58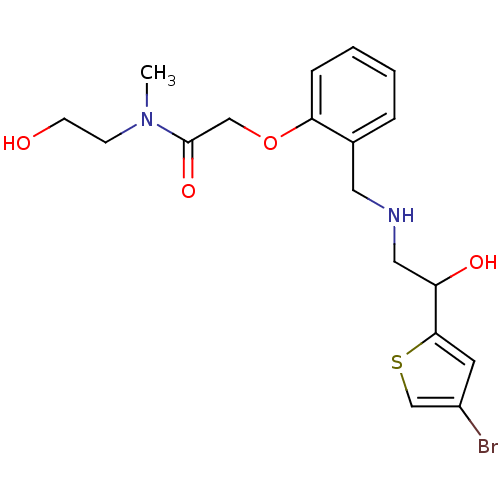 BDBM18107 ethanolamine, 52 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylacetamide
BDBM18107 ethanolamine, 52 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylacetamide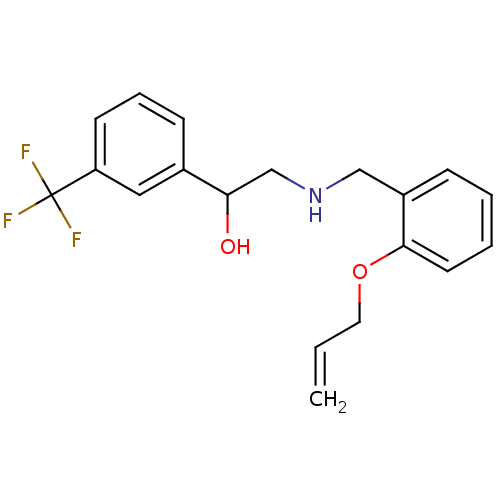 ethanolamine, 12 BDBM18080 2-({[2-(prop-2-en-1-yloxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol
ethanolamine, 12 BDBM18080 2-({[2-(prop-2-en-1-yloxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol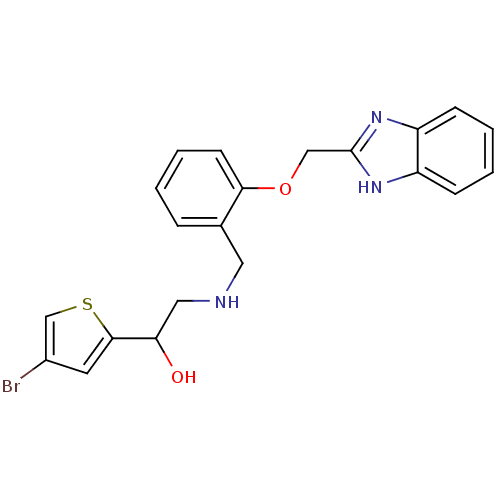 BDBM18103 ethanolamine, 48 2-({[2-(1H-1,3-benzodiazol-2-ylmethoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol
BDBM18103 ethanolamine, 48 2-({[2-(1H-1,3-benzodiazol-2-ylmethoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol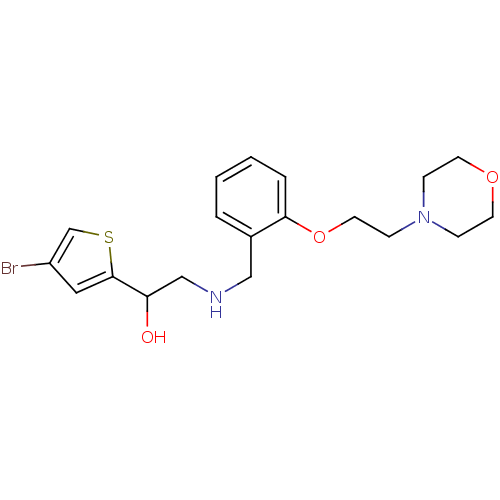 ethanolamine, 41 1-(4-bromothiophen-2-yl)-2-[({2-[2-(morpholin-4-yl)ethoxy]phenyl}methyl)amino]ethan-1-ol BDBM18096
ethanolamine, 41 1-(4-bromothiophen-2-yl)-2-[({2-[2-(morpholin-4-yl)ethoxy]phenyl}methyl)amino]ethan-1-ol BDBM18096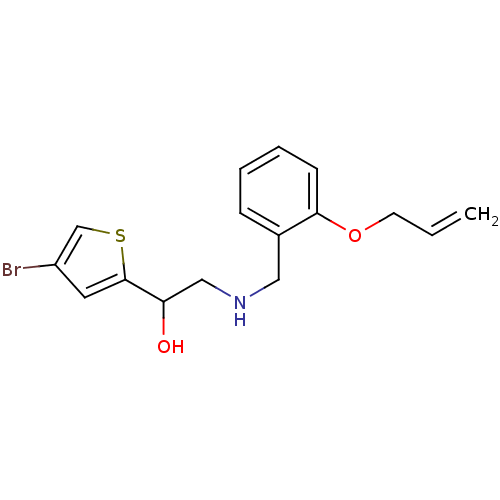 ethanolamine, 42 BDBM18097 1-(4-bromothiophen-2-yl)-2-({[2-(prop-2-en-1-yloxy)phenyl]methyl}amino)ethan-1-ol
ethanolamine, 42 BDBM18097 1-(4-bromothiophen-2-yl)-2-({[2-(prop-2-en-1-yloxy)phenyl]methyl}amino)ethan-1-ol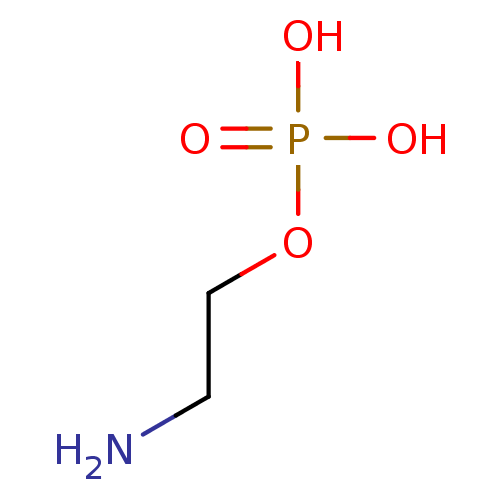 2-amino-ethanol dihydrogen phosphate O-phosphocolamine ethanolamine O-phosphate OPE phosphonoethanolamine monoaminoethyl phosphate colaminphosphoric acid 2-amino-ethanol phosphate PEA colamine phosphate mono(2-aminoethyl) phosphate colamine phosphoric acid 2-aminoethyl dihydrogen phosphate phosphoric acid 2-aminoethyl phenyl ester CHEMBL146972 BDBM50281572 EAP ethanolamine acid phosphate pEtN O-phosphoethanolamine
2-amino-ethanol dihydrogen phosphate O-phosphocolamine ethanolamine O-phosphate OPE phosphonoethanolamine monoaminoethyl phosphate colaminphosphoric acid 2-amino-ethanol phosphate PEA colamine phosphate mono(2-aminoethyl) phosphate colamine phosphoric acid 2-aminoethyl dihydrogen phosphate phosphoric acid 2-aminoethyl phenyl ester CHEMBL146972 BDBM50281572 EAP ethanolamine acid phosphate pEtN O-phosphoethanolamine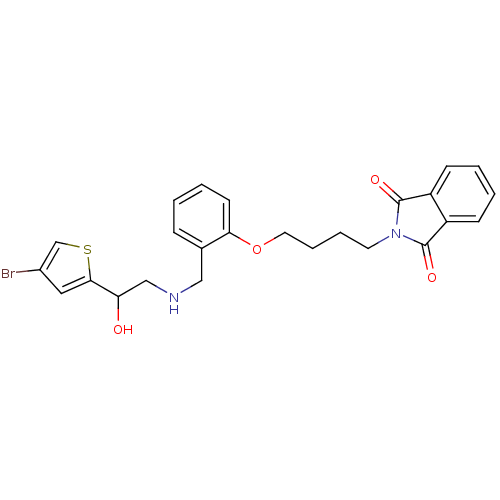 BDBM18115 ethanolamine, 60 2-{4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]butyl}-2,3-dihydro-1H-isoindole-1,3-dione
BDBM18115 ethanolamine, 60 2-{4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]butyl}-2,3-dihydro-1H-isoindole-1,3-dione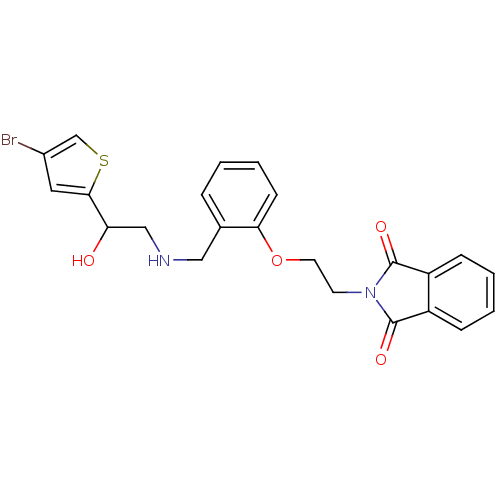 ethanolamine, 40 BDBM18095 2-{2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethyl}-2,3-dihydro-1H-isoindole-1,3-dione
ethanolamine, 40 BDBM18095 2-{2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethyl}-2,3-dihydro-1H-isoindole-1,3-dione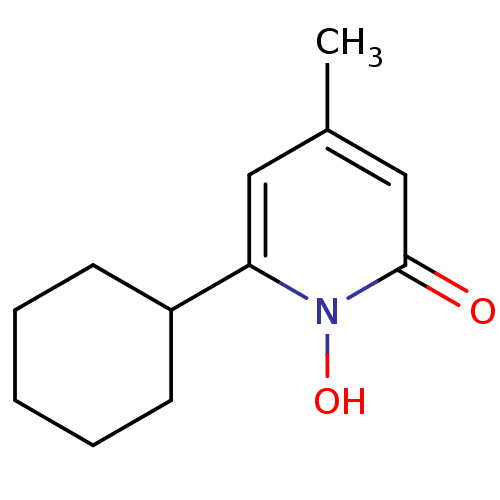 MLS002153867 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methyl-2-pyridinone BDBM66087 cid_38911 2-azanylethanol;6-cyclohexyl-4-methyl-1-oxidanyl-pyridin-2-one Ciclopirox Ciclopirox ethanolamine SMR001233223 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methyl-2-pyridone 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methylpyridin-2-one
MLS002153867 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methyl-2-pyridinone BDBM66087 cid_38911 2-azanylethanol;6-cyclohexyl-4-methyl-1-oxidanyl-pyridin-2-one Ciclopirox Ciclopirox ethanolamine SMR001233223 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methyl-2-pyridone 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methylpyridin-2-one
- Jarvest, RL; Erskine, SG; Forrest, AK; Fosberry, AP; Hibbs, MJ; Jones, JJ; Oamppound39Hanlon, PJ; Sheppard, RJ; Worby, A Discovery and optimisation of potent, selective, ethanolamine inhibitors of bacterial phenylalanyl tRNA synthetase. Bioorg Med Chem Lett 15: 2305-9 (2005)
- Gilmore, JL; Sheppeck, JE; Watterson, SH; Haque, L; Mukhopadhyay, P; Tebben, AJ; Galella, MA; Shen, DR; Yarde, M; Cvijic, ME; Borowski, V; Gillooly, K; Taylor, T; McIntyre, KW; Warrack, B; Levesque, PC; Li, JP; Cornelius, G; D'Arienzo, C; Marino, A; Balimane, P; Salter-Cid, L; Barrish, JC; Pitts, WJ; Carter, PH; Xie, J; Dyckman, AJ Discovery and Structure-Activity Relationship (SAR) of a Series of Ethanolamine-Based Direct-Acting Agonists of Sphingosine-1-phosphate (S1P1). J Med Chem 59: 6248-64 (2016)
- ChEMBL_643589 (CHEMBL1212453) Inhibition of mouse brain FAAH assessed as conversion of [3H-ethanolamine]AEA to [3H]ethanolamine by scintillation counting
- ChEMBL_1572612 (CHEMBL3803145) Inhibition of FAAH in rat brain membrane using anandamide[ethanolamine-3H] as substrate assessed as reduction in [3H]-ethanolamine production incubated for 30 mins by scintillation counting method
- ChEMBL_1776175 (CHEMBL4233167) Inhibition of ACAT1 in human monocytes-derived macrophages reduction in cholesteryl oleate formation preincubated for 2 hrs followed by [14C]-oleic acid/BSA/sodium oleate addition measured after 2 hrs by TLC assay
- ChEMBL_2077560 (CHEMBL4733351) Inhibition of FAAH in human U937 cells assessed as reduction in [3H]ethanolamine formation using [ethanolamine-1-3H]AEA as substrate incubated for 15 mins by liquid scintillation spectroscopy
- ChEBML_154781 Inhibition of pancreatic cholesterol esterase using p-nitrophenylacetate or cholesteryl [1-14C]-oleate as substrates
- ChEMBL_1776208 (CHEMBL4233200) Inhibition of ACAT1 in human monocytes-derived macrophages reduction in cholesteryl oleate formation preincubated for 2 hrs followed by [14C]-oleic acid/sodium oleate addition measured after 2 hrs in absence of albumin by TLC assay
- ChEMBL_1566101 (CHEMBL3791985) Inhibition of FAAH in rat brain membranes using using N-arachidonoyl-[14C]-ethanolamine as substrate assessed as reduction in [14C]ethanolamine production incubated for 30 mins by scintillation counting method
- ChEMBL_582084 (CHEMBL1060803) Inhibition of human SCD1 expressed in HEK293A cells assessed as conversion of [14C]stearate to [14C]oleate
- FAAH Carbon Filtration Assay [3H]anandamide was incubated with membranes to produce radiolabeled ethanolamine and unlabeled arachidonic acid. Charcoal selectively binds anandamide and arachidonic acid, but not ethanolamine. Uncleaved [3H] anandamide and the unlabeled arachidonic acid are absorbed by the charcoal. In contrast, the labeled [3H] ethanolamine flows through the charcoal minicolumns. The plates can then be read on a Hewlett-Packard TopCount.
- ChEMBL_590923 (CHEMBL1045641) Displacement of [3H]ethanolamine from human recombinant FAAH by liquid scintillation counting
- ChEMBL_428853 (CHEMBL917129) Inhibition of human group IB PLA2 in [3H]oleate-labeled Escherichia coli membrane by radiometric assay
- ChEMBL_428854 (CHEMBL917130) Inhibition of human group IIA PLA2 in [3H]oleate-labeled Escherichia coli membrane by radiometric assay
- ChEMBL_428855 (CHEMBL917131) Inhibition of human group V PLA2 in [3H]oleate-labeled Escherichia coli membrane by radiometric assay
- ChEMBL_428856 (CHEMBL917132) Inhibition of human group X PLA2 in [3H]oleate-labeled Escherichia coli membrane by radiometric assay
- ChEMBL_577393 (CHEMBL1056245) Inhibition of stearoyl-CoA desaturase 1 in mouse microsome assessed as conversion of [14C]stearate to [14C]oleate
- ChEMBL_577394 (CHEMBL1056246) Inhibition of stearoyl-CoA desaturase 1 in human microsome assessed as conversion of [14C]stearate to [14C]oleate
- ChEMBL_582072 (CHEMBL1060791) Inhibition of SCD1 in mouse microsomes assessed as conversion of [14C]stearate to [14C]oleate after 60 mins
- ChEMBL_582073 (CHEMBL1060792) Inhibition of SCD1 in human microsomes assessed as conversion of [14C]stearate to [14C]oleate after 60 mins
- ChEMBL_606060 (CHEMBL1071959) Inhibition of human SCD1 expressed in human 293A cells assessed as conversion of [14C]stearate to [14C]oleate
- ChEBML_156040 Inhibition of secretory PLA2 activity in bee venom enzyme using [3H]oleate labelled membranes at a concentration of 10 uM
- ChEMBL_156348 (CHEMBL761679) Inhibition of secretory PLA2 activity in porcine pancreas using [3H]-oleate labelled membranes at a concentration of 10 uM
- ChEMBL_1646376 (CHEMBL3995432) Inhibition of human pancreatic lipase using 4-methylumbelliferyl oleate as substrate after 30 mins by fluorometric method
- ChEMBL_1566100 (CHEMBL3791984) Inhibition of recombinant C-terminal His-tagged human FAAH expressed in sf21 cells using N-arachidonoyl-[14C]-ethanolamine as substrate assessed as reduction in [14C]ethanolamine production incubated for 30 mins by scintillation counting method
- ChEBML_156195 Inhibition of secretory PLA2 activity in human recombinant synovial enzyme using [3H]oleate labelled membranes at a concentration of 10 uM
- ChEMBL_156040 (CHEMBL764348) Inhibition of secretory PLA2 activity in bee venom enzyme using [3H]oleate labelled membranes at a concentration of 10 uM
- ChEMBL_155887 (CHEMBL769604) Inhibition of secretory PLA2 activity in Naja naja venom enzyme using [3H]oleate labelled membranes at a concentration of 10 uM
- ChEMBL_2033827 (CHEMBL4687985) Inhibition of CETP in 95% human serum assessed as inhibition of transfer of [3H] cholesteryl oleate or [3H] triolein between LDL and HDL
- ChEMBL_740688 (CHEMBL1763750) Inhibition of FAAH in human T84 cell assessed as conversion of [3H]AEA to [3H]ethanolamine by scintillation counting
- ChEMBL_740702 (CHEMBL1763764) Inhibition of FAAH in rat RBL cell assessed as conversion of [3H]AEA to [3H]ethanolamine by scintillation counting
- ChEMBL_606059 (CHEMBL1071958) Inhibition of SCD1 in mouse microsomal liver S9 microsomal fraction assessed as conversion of [14C]stearate to [14C]oleate after 60 mins
- ChEMBL_88053 (CHEMBL693216) Inhibitory activity against Human Hormone-sensitive lipase (HSL) expressed in the baculovirus was determined by HSL [14 C] cholesteryl oleate assay
- ChEMBL_1464456 (CHEMBL3405398) Inhibition of human FAAH using [ethanolamine 1-3H] substrate assessed as radioactivity by liquid scintillation counting analysis
- ChEMBL_606058 (CHEMBL1071957) Inhibition of SCD1 in human 293A cells assessed as conversion of [14C]stearate to [14C]oleate after 60 mins in presence of S9 microsomal fraction
- ChEMBL_606061 (CHEMBL1071960) Inhibition of SCD1 in human liver S9 microsomal fraction assessed as conversion of [14C]stearate to [14C]oleate after 60 mins at 0.4 uM
- ChEMBL_606657 (CHEMBL1071884) Inhibition of Scd1 in mouse liver microsomes assessed as conversion of [14C]stearate to [14C]oleate pretreated for 10 mins measured after 60 mins
- ChEBML_1558646 Inhibition of rat brain FAAH assessed as hydrolysis of [14C]AEA to [14C]Ethanolamine incubated for 30 mins by scintillation counting method
- ChEMBL_606658 (CHEMBL1071885) Inhibition of human Scd1 expressed in HEK293A cells assessed as conversion of [14C]stearate to [14C]oleate pretreated for 30 mins measured after 4 hrs
- ChEMBL_606659 (CHEMBL1071886) Inhibition of Scd1 in human HEK293A cell microsomes assessed as conversion of [14C]stearate to [14C]oleate pretreated for 10 mins measured after 60 mins
- ChEMBL_1558646 (CHEMBL3771523) Inhibition of rat brain FAAH assessed as hydrolysis of [14C]AEA to [14C]Ethanolamine incubated for 30 mins by scintillation counting method
- ChEMBL_2029682 (CHEMBL4683840) Inhibition of FAAH in OF1 albino mouse brain homogenate assessed as reduction in [3H]ethanolamine production using [3H]anandamide as substrate
- ChEMBL_789477 (CHEMBL1924504) Inhibition of human recombinant FAAH assessed as conversion of [3H]AEA to [3H]ethanolamine after 10 mins by liquid scintillation counting
- ChEMBL_856312 (CHEMBL2160519) Inhibition of human recombinant FAAH assessed as conversion of [3H]AEA to [3H]ethanolamine after 10 mins by liquid scintillation counter
- ChEMBL_2033826 (CHEMBL4687984) Inhibition of recombinant CETP (unknown origin) assessed as inhibition of transfer of [3H] cholesteryl oleate or [3H] triolein between exogenous LDL and HDL in 2% human serum
- ChEMBL_804415 (CHEMBL1954842) Inhibition of human endothelial lipase expressed using recombinant adenovirus using glycerol-tri[9,10(n)-3H]oleate after 1 hr by vesicle assay
- ChEMBL_804416 (CHEMBL1954843) Inhibition of human lipoprotein lipase expressed using recombinant adenovirus using glycerol-tri[9,10(n)-3H]oleate after 1 hr by vesicle assay
- ChEMBL_968124 (CHEMBL2398982) Non-competitive inhibition of microsomal ACAT in human MDM using [14C]oleoyl-CoA as substrate assessed as formation of cholesteryl [14C]-oleate after 24 hrs
- ChEMBL_1589177 (CHEMBL3829832) Inhibition of FAAH in rat brain membrane using N-arachidonoyl-[14C]-ethanolamine as substrate incubated for 30 mins by scintillation counting method
- ChEMBL_956683 (CHEMBL2379080) Inhibition of FAAH in rat brain membranes assessed as [14C]AEA hydrolysis to [14C]Ethanolamine after 30 mins by scintillation counting analysis
- ChEMBL_1808007 (CHEMBL4307366) Inhibition of DGAT1 in mouse C2C12 cells assessed as reduction in intracellular triglyceride production incubated for 2 hrs in presence of BSA-complexed oleate by LC/MS/MS analysis
- ChEMBL_1692729 (CHEMBL4043619) Inhibition of FAAH in human U937 cells using [ethanolamine-1-3H]AEA as substrate after 15 mins by liquid scintillation counting method
- ChEMBL_1921762 (CHEMBL4424607) Inhibition of FAAH in human U937 cell homogenates using [ethanolamine-1-3H]AEA as substrate after 15 mins by liquid scintillation spectroscopy
- ChEMBL_1577866 (CHEMBL3808209) Inhibition of CETP in 95% human serum assessed as suppression of transfer of [3H]-cholesterol oleate/[3H]-triolein from LDL to HDL after 1 hr by liquid scintillation assay
- ChEMBL_968111 (CHEMBL2398969) Non-competitive inhibition of human microsomal ACAT2 overexpressed in CHO cells using [14C]oleoyl-CoA as substrate assessed as formation of cholesteryl [14C]-oleate after 24 hrs
- ChEMBL_968112 (CHEMBL2398970) Non-competitive inhibition of human microsomal ACAT1 overexpressed in CHO cells using [14C]oleoyl-CoA as substrate assessed as formation of cholesteryl [14C]-oleate after 24 hrs
- ChEMBL_608772 (CHEMBL1068364) Inhibition of human recombinant NAAA expressed in human HEK293 cells assessed as conversion of [1,2-14C]palmitoylethanolamine to [1,2-14C]ethanolamine by liquid scintillation counting
- ChEMBL_933546 (CHEMBL2319708) Inhibition of FAAH in rat brain homogenates using [14C]anandamide as substrate assessed as formation of [14C]ethanolamine after 30 mins by scintillation counting analysis
- ChEMBL_969749 (CHEMBL2404091) Inhibition of Wistar rat brain FAAH using [3H]-ethanolamine as substrate preincubated for 20 mins followed by substrate addition by liquid scintillation counting analysis
- ChEMBL_1992822 (CHEMBL4626557) Inhibition of SCD in human NCI-H2122 cells expressing CYP4F11 assessed as reduction in cell viability incubated for 96 hrs in presence of oleate by cell titer Glo reagent based assay
- ChEMBL_2133982 (CHEMBL4843592) Inhibition of recombinant CETP (unknown origin) assessed as inhibition of transfer of [3H]cholesteryl oleate or [3H]triolein between exogenous [3H]LDL in 95% human serum by liquid scintillation analysis
- ChEMBL_2134010 (CHEMBL4843620) Inhibition of recombinant CETP (unknown origin) assessed as inhibition of transfer of [3H]cholesteryl oleate or [3H]triolein using exogenous LDL and HDL in 2% human serum by liquid scintillation analysis
- ChEMBL_2261405 (CHEMBL5216416) Displacement of [14C]-ethanolamine from FAAH derived from rat brain membrane fraction preincubated for 20 mins followed by susbstrate addition by scintillation counting method
- ChEMBL_939360 (CHEMBL2327495) Apparent inhibition of human FAAH expressed in CHO-K1 cells using ethanolamine 1-3[H] as substrate after 30 mins by liquid scintillation counting
- ChEBML_1692736 Inhibition of FAAH in human U937 cells assessed as decrease in [ethanolamine-1-3H]AEA uptake preincubated for 2 mins followed by AEA addition after 5 mins by liquid scintillation counting method
- ChEMBL_1589176 (CHEMBL3829831) Inhibition of FAAH in rat brain membrane using N-arachidonoyl-[14C]-ethanolamine as substrate preincubated for 20 mins followed by substrate addition measured after 30 mins by scintillation counting method
- ChEMBL_2056356 (CHEMBL4711357) Inhibition of FAAH in human U937 cells using [ethanolamine-1-3H]AEA as substrate preincubated for 15 mins followed by substrate addition and measured after 15 mins by liquid scintillation spectroscopy
- FAAH Inhibition Assay FAAH Inhibition Assay [3H]Ethanolamine produced from [3H]AEA hydrolysis was used to calculate FAAH activity and was measured by scintillation counting of the aqueous phase after extraction of the incubation mixture. Data were expressed as the percentage of [3H]-ethanolamine formed versus vehicle, after subtraction of the background radioactivity determined in the presence of arachidonyl trifluoromethyl ketone (ATFMK), an inhibitor of FAAH. IC50 values were determined using a four-parameter logistic equation for dose-response curves.
- ChEMBL_1692736 (CHEMBL4043626) Inhibition of FAAH in human U937 cells assessed as decrease in [ethanolamine-1-3H]AEA uptake preincubated for 2 mins followed by AEA addition after 5 mins by liquid scintillation counting method
- ChEMBL_1802959 (CHEMBL4275251) Inhibition of FAAH in human U937 cells using [ethanolamine-1-3H]AEA as substrate pretreated for 30 mins followed by substrate addition and measured after 15 mins by liquid scintillation counting method
- ChEMBL_858269 (CHEMBL2168075) Inhibition of mouse FAAH isolated from brain homogenate using [3H-ethanolamine]AEA as substrate incubated for 20 mins prior to substrate addition measured after 15 mins by beta counting analysis
- ChEMBL_1589181 (CHEMBL3829836) Inhibition of FAAH in rat brain membrane at pH 9 using N-arachidonoyl-[14C]-ethanolamine as substrate preincubated for 10 mins followed by substrate addition measured after 30 mins by scintillation counting method
- ChEMBL_1589182 (CHEMBL3829837) Inhibition of FAAH in rat brain membrane at pH 7.4 using N-arachidonoyl-[14C]-ethanolamine as substrate preincubated for 10 mins followed by substrate addition measured after 30 mins by scintillation counting method
- ChEMBL_1739438 (CHEMBL4155188) Inhibition of FAAH in human U937 cells using AEA containing [ethanolamine-1-3H]AEA as substrate preincubated for 30 mins followed by substrate addition and measured after 15 mins by liquid scintillation spectroscopic method
- ChEMBL_1746160 (CHEMBL4180670) Inhibition of FAAH in human U937 cell homogenate assessed as decrease in [ethanolamine-1-3H]AEA levels preincubated for 30 mins followed by AEA addition measured after 15 mins by liquid scintillation counting method
- ChEMBL_1746161 (CHEMBL4180671) Inhibition of MAGL in human U937 cell homogenate assessed as decrease in [ethanolamine-1-3H]AEA levels preincubated for 30 mins followed by AEA addition measured after 15 mins by liquid scintillation counting method
- ChEMBL_2131321 (CHEMBL4840836) Inhibition of FAAH (unknown origin) in human U-937 cells using [ethanolamine-1-3H]AEA as substrate preincubated for 30 mins followed by substrate addition measured after 15 mins by liquid scintillation spectroscopy
- ChEMBL_2131322 (CHEMBL4840837) Inhibition of MAGL (unknown origin) in human U-937 cells using [ethanolamine-1-3H]AEA as substrate preincubated for 30 mins followed by substrate addition measured after 15 mins by liquid scintillation spectroscopy
- ChEMBL_934690 (CHEMBL2318384) Inhibition of FAAH-mediated [3H]anandamide hydrolysis in rat brain homogenate assessed as [3H]ethanolamine production incubated for 45 mins prior to substrate addition measured after 10 mins by liquid scintillation counting analysis
- ChEMBL_934692 (CHEMBL2318386) Inhibition of FAAH-mediated [3H]anandamide hydrolysis in rat brain homogenate assessed as [3H]ethanolamine production incubated for 90 mins prior to substrate addition measured after 10 mins by liquid scintillation counting analysis
- FAAH Inhibition Assay [3H]Ethanolamine produced from [3H]AEA hydrolysis was used to calculate FAAH activity and was measured by scintillation counting of the aqueous phase after extraction of the incubation mixture. Data are expressed as the concentration exerting a half maximal inhibition (IC50).
- ChEMBL_2029680 (CHEMBL4683838) Inhibition of FAAH in Sprague-Dawley rat brain homogenate using arachidonoyl-ethanolamide and anandamide [ethanolamine-1-3H] as substrate pre-incubated for 10 mins followed by substrate addition and measured after 30 mins by liquid scintillation counting method
- FAAH Inhibition Assay Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3H], 60 Cimmol-1, from American Radiolabeled Chemicals, St. Louis, USA) as a substrate.
- In Vitro Pancreatic Lipase Assay Various concentrations of test compound (5, 10, 25, 50 and 100 mM) were dissolved in DMSO (final culture concentration 0.1%) and 4-MU oleate and lipase were diluted in 0.1M McIlvane buffer containing 0.1M citric acid-Na2HPO4 (pH 7.4). The mixedsolution that composed of 100 mL of 0.1mM 4-MU oleate, 40 mL of the above-mentioned buffer and 10 mL of test samples was prepared before adding lipase. The enzymatic reaction was started by adding 0.05mL of 1 U/mL pancreatic lipase and incubation was carried out for 20 min at 37 �C. The amount of 4-methylumbelliferon released by the enzyme was monitored at an emission of 450 nm and excitation wavelength of 320 nm using a fluorescence spectrophotometer.
- FAAH Assay Enzyme activity was demonstrated in a radioenzymatic test based on measuring the product of hydrolysis (ethanolamine [3]H) of anandamide [ethanolamine 1-.sup.3H] (American Radiolabeled Chemicals; 1 mCi/ml) with FAAH (Life Sciences (1995), 56, 1999-2005 and Journal of Pharmacology and Experimented Therapeutics (1997), 283, 729-734), Analytical. Biochemistry (2003), 318, 270-5. In addition, routine assays were performed monitoring hydrolysis of arachidonyl-7-amino-4-methylcoumarin amide (AAMCA) by following increase in fluorescence upon release of 7-amino 4-methyl coumarin (λEX=355 nm, (λEM=460 nm). Analytical. Biochemistry (2005), 343, 143-51. Assays are performed on either cell lysate fractions prepared as described or in whole cell format employing either the fluorescent substrate AAMCA (Cayman chemical, Ann Arbor, Mich.,) or 3H-anandamide ([ETHANOLAMINE-1-3H] American Radiolabeled Chemicals; 1 mCi/ml). The cell lysate assay is performed in black PerkinElmer OptiPlates-384F.
- FAAH Inhibition Assay The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethanolamine produced from [14C]AEA hydrolysis was used to calculate FAAH activity and was measured by scintillation counting of the aqueous phase after extraction of the incubation mixture. Data are expressed as the concentration exerting a half maximal inhibition (IC50).
- FAAH Inhibition Assay The effect of the test compounds on the enzymatic hydrolysis of [3H]anandamide was evaluated by using rat brain homogenate preparations. [3H]Ethanolamine produced from [3H]AEA hydrolysis was used to calculate FAAH activity and was measured by scintillation counting of the aqueous phase after extraction of the incubation mixture. Data are expressed as the concentration exerting a half maximal inhibition (IC50).
- ACAT-1 Enzyme Inhibition Assay The assay mixture contains test compound, membrane protein, cholesterol solubilized in Triton WR-1339, and glutathione in a 96-deep well plate. The reaction was started by adding substrate [14C]oleoyl coenzyme A and incubated. After the reaction was stopped, radioactive cholesteryl oleate product was separated into the organic heptane phase. ACAT-1 activity was quantified by counting aliquots of the upper heptane layer by liquid scintillography.
- In Vitro Radioactive (RTA) Assay An in vitro assay for determining ICso's to identify compounds that inhibit CETP transfer activity is performed based on a modification of a published method (Morton and Zilversmit, (1981) A plasma inhibitor of triglyceride and cholesteryl ester transfer activities, J. Biol. Chem. 256(23), 1 1992-1 1995). The ability of inhibitors to alter CETP activity is performed using two different assays: one using recombinant CETP and one using an endogenous plasma source of CETP. Both assays measure the transfer of [3H] cholesteryl oleate or [3H] triolein from exogenous LDL to HDL.Radiolabeled donor particles are generated by first combining 100 ul of 200 uM butylated hydroxyl toluene in CHC13, 216 of 21.57 mM DOPC in EtOH, and either 500 uCi [3H]-triolein (Perkin Elmer #NET-431) or 500 uCi [3H]-cholesteryl oleate (GE #TRK886) in a glass tube. Reagents are mixed, dried under nitrogen, and then resuspended in 2 mL of 50 mM Tris, 27 uM EDTA.
- High Throughput FAAH Inhibition Assays The assays for compounds described herein are amenable to high throughput screening. Preferred assays thus detect binding of the inhibitor to FAAH or the release of a reaction product (e.g., fatty acid amide or ethanolamine) produced by the hydrolysis of a substrate such as oleoylethanolamide or anandamide. The substrate may be labeled to facilitate detection of the released reaction products. High throughput assays for the presence, absence, or quantification of particular reaction products are well known to those of skill in the art. Thus, for example, U.S. Pat. No. 5,559,410 discloses high throughput screening methods for proteins, and U.S. Pat. Nos. 5,576,220 and 5,541,061 disclose high throughput methods of screening for ligand/antibody binding.
- FAAH Assay 1 T84 frozen cell pellets or transfected SK-N-MC cells (contents of 1�15 cm culture dishes) were homogenized in 50 mL of FAAH assay buffer (125 mM Tris, 1 mM EDTA, 0.2% Glycerol, 0.02% Triton X-100, 0.4 mM Hepes, pH 9). The assay mixture consisted of 50 μL of the cell homogenate, 10 μL of the test compound, and 40 μL of anandamide [1-3H-ethanolamine] (3H-AEA, Perkin-Elmer, 10.3 Ci/mmol), which was added last, for a final tracer concentration of 80 nM. The reaction mixture was incubated at rt for 1 h. During the incubation, 96-well Multiscreen filter plates (catalog number MAFCNOB50; Millipore, Bedford, Mass., USA) were loaded with 25 μL of activated charcoal (Multiscreen column loader, catalog number MACL09625, Millipore) and washed once with 100 μL of MeOH. Also during the incubation, 96-well DYNEX MicroLite plates (catalog number NL510410) were loaded with 100 μL of MicroScint40 (catalog number 6013641, Packard Bioscience, Meriden, Conn., USA). After the 1 h incubation, 60 μL of the reaction mixture were transferred to the charcoal plates, which were then assembled on top of the DYNEX plates using Centrifuge Alignment Frames (catalog number MACF09604, Millipore). The unbound labeled ethanolamine was centrifuged through to the bottom plate (5 min at 2000 rpm), which was preloaded with the scintillant, as described above. The plates were sealed and left at rt for 1 h before counting on a Hewlett Packard TopCount.
- FAAH Assay 2 Transfected SK-N-MC cells (contents of 1�15 cm culture dishes) were homogenized in 50 mL of FAAH assay buffer (125 mM Tris, 1 mM EDTA, 0.2% Glycerol, 0.02% Triton X-100, 0.4 mM Hepes, pH 9). The assay mixture consisted of 50 μL of the cell homogenate, 10 μL of the test compound, and 40 μL of anandamide [1-3H-ethanolamine] (3H-AEA, Perkin-Elmer, 10.3 Ci/mmol), which was added last, for a final tracer concentration of 80 nM. The reaction mixture was incubated at rt for 1 h. During the incubation, 96-well Multiscreen filter plates (catalog number MAFCNOB50; Millipore, Bedford, Mass., USA) were loaded with 25 μg of activated charcoal (Multiscreen column loader, catalog number MACL09625, Millipore) and washed once with 100 μL of MeOH. Also during the incubation, 96-well DYNEX MicroLite plates (catalog number NL510410) were loaded with 100 μL of MicroScint40 (catalog number 6013641, Packard Bioscience, Meriden, Conn., USA). After the 1 h incubation, 60 μL of the reaction mixture were transferred to the charcoal plates, which were then assembled on top of the DYNEX plates using Centrifuge Alignment Frames (catalog number MACF09604, Millipore). The unbound labeled ethanolamine was centrifuged through to the bottom plate (5 min at 2000 rpm), which was preloaded with the scintillant, as described above. The plates were sealed and left at rt for 1 h before counting on a Hewlett Packard TopCount.
- FAAH Activity Hydrolysis of [3H]-AEA by FAAH was determined as previously described in cell homogenates of U937 cells (0.18 mg protein) (Omeir et al., 1999, Biochem Biophys Res Commun, 264, 316-20; Mor et al., 2004, J Med Chem, 47, 4998-5008). Protein amounts of cell homogenates corresponded to 0.5�106 cells (U937), to assure best possible comparability of IC50 values as used for the AEA cellular uptake assays. URB597 was used as positive control. Protein quantification was performed using a BCA assay (Thermo Scientific). Enzyme activity was assessed by addition of vehicle or compounds in 10 μL DMSO to 490 μL homogenate in 10 mM Tris HCl, 1 mM EDTA, 0.1% (w/v) BSA fatty acid free, pH=8 and incubation for 15 min at 37� C. After, a mixture of AEA plus [ethanolamine-1-3H]-AEA (0.5 nM) at final 100 nM was added to the homogenates and incubated for 15 min at 37� C. The reaction was stopped by addition of 1 mL ice-cold CHCl3:MeOH (1:1) followed by vigorous vortexing. Phase separation was achieved by centrifugation at 10′000�g at 4� C. for 10 min. Radioactivity of the separated aqueous phase (upper phase) containing [3H-ethanolamine] or [3H-glycerol] was measured by liquid scintillation counting on a Tri-Carb 2100 TR liquid scintillation analyzer after addition of 3.5 mL Ultima Gold scintillation cocktail (PerkinElmer Life Sciences). Results are expressed as hydrolysis of tritium substrate in percent of vehicle treated control. IC50 values were calculated by GraphPad . Data are reported as means of n=3 independent experiments performed in triplicates.
- hFAP Binding Assay Table 1: Compounds were tested in a direct binding assay using 8K surface plasmon resonance biosensor (GE Healthcare) at 20� C. Immobilization of hFAP (M39-A757) on a CMD200M sensor chip (Xantec) was performed using standard amine coupling procedure in immobilization buffer (10 mM HEPES, 150 mM NaCl, 0.05% Tween20, pH 7.4). The surface was washed with 10 mM NaOH, 1M NaCl before being activated with EDC/NHS (GE Healthcare), followed by immobilization of hFAP (in 10 mM Acetate pH 5.0). Finally, the surface was deactivated by ethanolamine. Immobilization levels of hFAP were around 4000-6000 RU. The reference spot was treated as described, omitting the injection of hFAP. Compound concentration series were injected over the immobilized protein in increasing concentrations (2-500 nM) using single cycle kinetics in running buffer (20 mM TRIS, 150 mM NaCl, 0.05% Tween20, 1% DMSO, pH 7.4).
- Secondary Binding Analysis by Surface Plasmon Resonance The His-tagged SARS-CoV-2 PLpro enzyme was initially prepared in phosphate buffer and diluted to 50 ug/mL with 10 mM sodium acetate (pH 5.5) and immobilized on a CM5 sensor chip by standard amine coupling with running buffer PBSP (10 mM phosphate, pH 7.4, 2.7 mM KCl, 137 mM NaCl, 0.05% Tween-20). The CM5 sensor chip surface was first activated by 1-ethyl-3-(3- (dimethylamino)propyl)carbodiimide hydrochloride (EDC)/N-hydroxysuccinimide (NHS) mixture using a Biacore 8K instrument (Cytiva). SARS-CoV-2 PLpro enzyme was immobilized to flow channels 1−4 followed by ethanolamine blocking on the unoccupied surface area, and immobilization levels for all four channels were similar at 12,000 RU. Each flow channel has its own reference channel, and blank immobilization using EDC/NHS and ethanolamine was done for all reference channels. Compound solutions with a series of increasing concentrations (0.049−30 uM at 2.5-fold dilution) were applied to all active and reference channels in SPR binding buffer (10 mM HEPES, pH 7.4, 150 mM NaCl, and 0.05% Tween-20, 0.5 mM TCEP, and 2% DMSO) at a 30 uL/min flow rate at 25 �C. The data were double referenced with a reference channel and zero concentration (2% DMSO) responses, and reference subtracted sensorgrams were fitted with 1:1 Langmuir kinetic model using a Biacore Insight evaluation software, producing two rate constants (ka and kd) (Figure S1). The equilibrium dissociation constants (KD) were determined from two rate constants (KD = kd/ ka). For steady-state affinity fittings, response units at each concentration were measured during the equilibration phase, and the KD values were determined by fitting the data to a single rectangular hyperbolic curve equation, where y is the response, ymax is the maximum response, and x is the compound concentration.
- Surface Plasmon Resonance (SPR) Assay To establish binding constants for the compounds/drugs to PAI-1, an indirect approach using surface plasmon resonance (SPR) was employed. Varying concentrations of each compound were preincubated with PAI-1 in solution and then passed over immobilized anhydrotrypsin (Molecular Innovations) using a Biacore 2000 optical biosensor, and the loss of PAI-1 binding to anhydrotrypsin was quantified. Bovine anhydrotrypsin was immobilized to CM5 SPR chips at the levels of ~2000 response unit (RU) in 10 mM sodium acetate, pH 5.0. The reference flow cell surface was left blank to serve as a control. Remaining binding sites were blocked by 1 M ethanolamine, pH 8.5. All binding reactions were carried out in assay buffer. PAI-1 at 2 nM was first incubated with the indicated concentrations of inhibitor in running buffer for at least 15 min at 23 C.The slope of the association phase of PAI-1 binding to an immobilized ligand (e.g., vitronectin or anhydrotrypsin) has been shown to have a linear relationship.
- Biological Assay Assay Method 1: T84 frozen pellets (contents of 1-4x15 cm culture dishes) were homogenized in 300 mL of FAAH assay buffer (125 mM Tris, 1 mM EDTA, 0.2% glycerol, 0.02% Triton X-100, 0.4 mM Hepes, pH 9). The assay mixture was prepared from 50 uL of the cell homogenate, 10 uL of the test compound, and 40 uL of anandamide [1-3H-ethanolamine] (3H-AEA; Perkin-Elmer, 10.3 Ci/mmol), which was added last, for a final tracer concentration of 200 nM. The reaction mixture was incubated at rt for 1 hour (h). During the incubation, 96-well Multiscreen filter plates (catalog number MAFCNOB50; Millipore, Bedford, Mass., USA) were loaded with 25 uL of activated charcoal (Multiscreen column loader, catalog number MACL09625, Millipore) and washed once with 100 uL of MeOH. Also during the incubation, 96-well DYNEX MicroLite plates (catalog number NL510410) were loaded with 100 uL of MicroScint40 (catalog number 6013641, Packard Bioscience, Meriden, Conn., USA).
- FAAH Assay T84 frozen cell pellets or transfected SK-N-MC cells (contents of 1x15 cm culture dishes) were homogenized in 50 mL of FAAH assay buffer (125 mM Tris, 1 mM EDTA, 0.2% Glycerol, 0.02% Triton X-100, 0.4 mM Hepes, pH 9). The assay mixture consisted of 50 uL of the cell homogenate, 10 uL of the test compound, and 40 pt of anandamide [1-3H-ethanolamine] (3H-AEA, Perkin-Elmer, 10.3 Ci/mmol), which was added last, for a final tracer concentration of 80 nM. The reaction mixture was incubated at rt for 1 h. During the incubation, 96-well Multiscreen filter plates (catalog number MAFCNOB50; Millipore, Bedford, Mass., USA) were loaded with 25 uL of activated charcoal (Multiscreen column loader, catalog number MACL09625, Millipore) and washed once with 100 uL of MeOH. Also during the incubation, 96-well DYNEX MicroLite plates (catalog number NL510410) were loaded with 100 uL of MicroScint40 (catalog number 6013641, Packard Bioscience, Meriden, Conn., USA).
- FAAH Assay T84 frozen cell pellets or transfected SK-N-MC cells (contents of 1x15 cm culture dishes) were homogenized in 50 mL of FAAH assay buffer (125 mM Tris, 1 mM EDTA, 0.2% Glycerol, 0.02% Triton X-100, 0.4 mM Hepes, pH 9). The assay mixture consisted of 50 uL of the cell homogenate, 10 uL of the test compound, and 40 uL of anandamide [1-3H-ethanolamine] (3H-AEA, Perkin-Elmer, 10.3 Ci/mmol), which was added last, for a final tracer concentration of 80 nM. The reaction mixture was incubated at rt for 1 h. During the incubation, 96-well Multiscreen filter plates (catalog number MAFCNOB50; Millipore, Bedford, Mass., USA) were loaded with 25 uL of activated charcoal (Multiscreen column loader, catalog number MACL09625, Millipore) and washed once with 100 uL of MeOH. Also during the incubation, 96-well DYNEX MicroLite plates (catalog number NL510410) were loaded with 100 uL of MicroScint40 (catalog number 6013641, Packard Bioscience, Meriden, Conn., USA).
- hFAP Binding Assay Table 1B: Compounds were tested in a direct binding assay using 8K surface plasmon resonance biosensor (GE Healthcare) at 20� C. Immobilization of hFAP (M39-A757) on a CMD200M sensor chip (Xantec) was performed using standard amine coupling procedure in immobilization buffer (10 mM HEPES, 150 mM NaCl, 0.05% Tween20, pH 7.4). The surface was washed with 10 mM NaOH, 1M NaCl before being activated with EDC/NHS (GE Healthcare), followed by immobilization of hFAP (in 10 mM Acetate pH 5.0). Finally, the surface was deactivated by ethanolamine. Immobilization levels of hFAP were around 4000-6000 RU. The reference spot was treated as described, omitting the injection of hFAP. Compound concentration series were injected over the immobilized protein in increasing concentrations (2-500 nM) using single cycle kinetics in running buffer (20 mM TRIS, 150 mM NaCl, 0.05% Tween20, 1% DMSO, pH 7.4). Interaction models were fitted globally to the experimental traces, enabling determination of kon, koff and Kd.
- SPR Assay The anti-His antibody (His Capture Kit, GE Healthcare) was immobilized on sensor chip CM5 (GE Healthcare), in a Biacore T200 or Biacore S200 instrument, to ∼10000 resonance units (RU) using a standard amine coupling procedure (Amine Coupling Kit, GE Healthcare). His-tagged aldose reductase (ALR2), at a concentration of 100 μg/mL, was captured on the anti-His antibody using dual injection consisting of a 3 min injection of protein followed by a 100 s activation with a mixture (1:1) of 0.2 M N-ethyl-N-(dimethylamino)propylcarbodiimide (EDC) and 50 mM N-hydroxysuccinimide (NHS), and final blocking with an injection of 1.0 Methanolamine (pH 8.5) for 100 s. Activation with EDC and NHS and blocking with ethanolamine stabilized the ALR2 capture and allowed the use of a single surface for the analysis of a number of compounds with no need for regeneration. The ALR2 capture level was ∼1000 RU, and all injections were performed at a flow rate of 10 μL/min. For the interaction analysis, the compound concentrations ranged from 3 to 10000 nM with a 5-fold dilution step, using a single-cycle kinetic approach with three or five concentrations or injections per cycle.
- LCAT Activity Measurement (In Vitro) A fraction composed of HDL3 (1.125
- Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org Lett 2006, 8 (10), 2023) was labeled with MR121 fluorophore (CAS 185308-24-1, 1-(3-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2′,3′-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid.Assay working solutions were made as follows:Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaCl2, 1 mM MgCl2, 0.01% Triton-X-100, pH 8.0;ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5� final concentration in assay buffer;
- Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org Lett 2006, 8 (10), 2023) was labeled with MR121 fluorophore (CAS 185308-24-1, 1-(3-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2′,3′-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid.Assay working solutions were made as follows:Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaCl2, 1 mM MgCl2, 0.01% Triton-X-100, pH 8.0;ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5� final concentration in assay buffer;MR121 substrate solution: MR121 substrate stock solution (800 μM MR121 substrate in DMSO), diluted to 2-5� final concentration in assay buffer.
- SPR Biosensor Assay An SPR assay was used to determine the binding affinities of compounds. Experiments were performed on a Biacore 3000 (Biacore, Uppsala, Sweden) instrument with CM5 research grade chips (Biacore, Uppsala, Sweden). Sensorgrams were recorded at a frequency of 2.5 Hz. EcIspF was immobilized using amine-coupling chemistry with readings between 4500 and 5500 response units were obtained. After immobilization, a 6 min injection at 5 uL min-1 of 1 M ethanolamine was used to quench excess active succinamide ester groups. SPR binding experiments with EcIspF were performed at 10 deg C in 50 mM sodium phosphate pH 7 and 2 mM MgCl2 at a flow rate of 30 uL min-1. The sensor surface was regenerated between experiments by applying the running buffer for 20 min to dissociate any ligand complex. This was followed by a further 20 min stabilization period. The 40 min total regeneration time between experiments helped to eliminate any carry-over of EcIspF bound to ligand. Each injection at a given concentration was repeated three times. Blank injections were included for each measurement series and subtracted from the data. The equilibrium (steady state) binding curves were analyzed by nonlinear regression and fit to a one-to-one Langmuir binding model. All ligands assessed by SPR were greater than 95% purity as established by high-performance liquid chromatography or combustion analysis.
- Surface Plasmon Resonance (SPR)Assay The interactions between compounds and ABAD were performed using the dual flow cell BIACORE 3000 instrument. Surface Plasmon Resonance (SPR) studies were performed on a BIACORE 3000 at 25° C. SPR binding experiments with ABAD were performed in phosphate-buffered saline (PBS, pH 7.4, 0.005% surfactant P20) as the running and the sample buffer. The surface of the sensor chip was first activated with mixtures of N-hydroxysuccinimide (NHS, 115 mg/ml) and N-(3-dimethyl-aminopropyl)-N'-ethyl-carbodiimide-hydrochloride (EDC, 750 mg/ml) for 7 minutes. ABAD was dissolved in PBS buffer (pH 5.0) at a concentration of 10 μg/ml. The protein was immobilized directly and covalently on the hydrophilic carboxymethylated dextran matrix of the CM5 sensor chip (BIACORE) by using the standard primary amine coupling reaction on a CM5 sensor chip according to standard procedures. After the immobilization of the protein, excess activated carboxylic acid groups were quenched with ethanolamine (1 M, pH 8.5). Special care was taken during injection of samples because of carryover effects. Special washing routines were used to clean the system before injection of new samples. In addition, predipping of needles was performed. The sample flow was 40 μl/minute in experiments performed for the determination of the kinetic and equilibrium constants. Regeneration of the surfaces between subsequent binding experiments was achieved by washing the surface extensively (>>1 hour) with buffer solution.
- Surface Plasmon Resonance (SPR) Assay Recombinant full-length hnRNPA1 (aa 1-320) and truncated versions of hnRNPA1, including the N-terminal RNA binding domain (aa 1-196), the middle region (aa 182-268), and the C-terminal region (aa 268-320), were individually covalently coupled to a dextran matrix (CM5 chip) using a Biacore T200 system (GE Healthcare), following the manufacturer's protocols. Immobilized proteins were activated with N-hydroxysuccinimide and 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide to form a carbodiamide linkage between carboxyl moieties on the dextran and primary amide groups on the protein. hnRNPA1 variants were dissolved in 10 mM acetate buffer (full-length, pH 4; N-terminal RNA binding domain and the C-terminal region, pH 4.5; middle region, pH 5) to a concentration of 20 μg/ml and flowed over the activated surface for 5 min with PBS. Unreacted groups were blocked with ethanolamine. PBS containing 0.05% (v/v) P20 (surfactant) and 5% DMSO was used for priming and conditioning the surface of the chips. Quercetin was serially diluted (from 200 to 3 μM) in PBS buffer and maintained in DMSO (5% (v/v)) (final concentrations from 10 to 0.15 μM). The baseline response was determined for 120 s (30 μl/min) before perfusing the immobilized proteins with the compounds to allow association to occur. Dissociation was then monitored over a period of an additional 120 s. The immobilized proteins were regenerated with 50 mM NaOH.
- In Vitro Rat FAAH Radiometric Assay Rat FAAH was prepared from male Sprague Dawley rat brains, homogenized in a potter in 20 mM of Tris HCl pH 7.4, 0.32 M sucrose.The radiometric assay used to measure FAAH activity was performed in Eppendorf tubes: 50 μg of total rat brain homogenate were pre-incubated in 445.5 μL of assay buffer (50 mM Tris-HCl pH 7.4, 0.05% Fatty acid-free-bovine serum albumin (BSA)) with 4.5 μL of inhibitor (at appropriate concentration in DMSO) or DMSO alone (to measure FAAH total activity) for 10 min at 37� C. The blank (no activity control) was prepared using 445.5 μL of assay buffer and 4.5 μL of DMSO without the 50 μg of total rat brain homogenate.After 10 min of pre-incubation with test compounds, the reaction was started by adding of 50 μL of substrate and incubating for 30 min at 37� C. The substrate was prepared in assay buffer in order to achieve the final concentration of 1 M arachidonoyl ethanolamide (Cayman Chemical N. 90050) and 0.6 nM anandamide [ethanolamine-1-3H] (American Radiolabeled Chemicals Inc, ART. 0626, Conc. 1 mCi/mL, S.A. 60 Ci/mmol). The reaction was stopped by adding cold 1:1 CHCl3/MeOH. After 10 min of centrifugation (845�g at 4� C.) 600 μL of aqueous phase were transferred into scintillation vials previously filled with 3 mL of scintillation fluid (Ultima Gold , Perkin Elmer Inc., Cat. 6013329). Radioactivity was measured by liquid scintillation counting (MicroBeta2 LumiJET Perkin Elmer Inc.).
- In Vitro Rat FAAH Radiometric Assay Rat FAAH was prepared from male Sprague Dawley rat brains, homogenized in a potter in 20 mM of Tris HCl pH 7.4, 0.32 M sucrose.The radiometric assay used to measure FAAH activity was performed in eppendorf tubes: 50 μg of total rat brain homogenate were pre-incubated in 445.5 μL of assay buffer (50 mM Tris-HCl pH7.4, 0.05% Fatty acid-free bovine serum albumin (BSA)) with 4.5 μL of inhibitor (at appropriate concentration in DMSO) or DMSO alone (to measure FAAH total activity) for 10 minutes at 37� C. The blank (no activity control) was prepared using 445.5 μL of assay buffer and 4.5 μL of DMSO without the 50 μg of total rat brain homogenate.After 10 minutes of pre-incubation with test compounds, the reaction was started by adding of 50 μL of substrate and incubating for 30 min at 37� C. The substrate was prepared in assay buffer in order to achieve the final concentration of 1 μM arachidonoyl ethanolamide (Cayman Chemical N. 90050) and 0.6 nM anandamide [ethanolamine-1-3H](American Radiolabeled Chemicals Inc., ART. 0626, conc. 1 mCi/mL, S.A. 60 Ci/mmol). The reaction was stopped by adding cold 1:1 CHCl3/MeOH. After 10 minutes of centrifugation (845�g at 4� C.) 600 μL of aqueous phase were transferred into scintillation vials previously filled with 3 mL of scintillation fluid (Ultima Gold, Perkin Elmer Inc., Cat. 6013329). Radioactivity was measured by liquid scintillation counting (MicroBeta2 LumiJET Perkin Elmer Inc.).
- Radiometric Assay Rat FAAH was prepared from male Sprague Dawley rat brains, homogenized in a potter in 20 mM of Tris HCl pH 7.4, 0.32 M sucrose.The radiometric assay used to measure FAAH activity was performed in Eppendorf tubes: 50 μg of total rat brain homogenate were pre-incubated in 445.5 μL of assay buffer (50 mM Tris-HCl pH 7.4, 0.05% Fatty acid-free-bovine serum albumin (BSA)) with 4.5 μL of inhibitor (at appropriate concentration in DMSO) or DMSO alone (to measure FAAH total activity) for 10 min at 37� C. The blank (no activity control) was prepared using 445.5 μL of assay buffer and 4.5 μL of DMSO without the 50 μg of total rat brain homogenate.After 10 min of pre-incubation with test compounds, the reaction was started by adding of 50 μL of substrate and incubating for 30 min at 37� C. The substrate was prepared in assay buffer in order to achieve the final concentration of 1 μM arachidonoyl ethanolamide (Cayman Chemical N. 90050) and 0.6 nM anandamide [ethanolamine-1-3H] (American Radiolabeled Chemicals Inc, ART. 0626, Conc. 1 mCi/mL, S.A. 60 Ci/mmol). The reaction was stopped by adding cold 1:1 CHCl3/MeOH. After 10 min of centrifugation (845�g at 4� C.) 600 μL of aqueous phase were transferred into scintillation vials previously filled with 3 mL of scintillation fluid (Ultima Gold , Perkin Elmer Inc., Cat. 6013329). Radioactivity was measured by liquid scintillation counting (MicroBeta2 LumiJET Perkin Elmer Inc.).
- SPR Binding Affinity Assay (Single Cycle) Single cycle kinetics (SCK): five increasing concentrations were injected successively without allowing for the dissociation of the protein-ligand complex. SPR experiments were performed at 25 °C using a Biacore T200 instrument (GE Healthcare). PBS (pH 7.4) supplemented with 0.05% Tween 20 was used as running buffer. Human FD and FB were immobilized covalently to a Series S Sensor Chip CM5 (GE Healthcare) at a flow rate of 10 μL/min using an amine-coupling protocol. Reagents for the immobilization were purchased from GE Healthcare (Amine Coupling Kit; BR-1000-50). The sensor-chip surface was activated by a 5 min injection of a 1:1 (v/v) mixture of a 100 mM N-hydroxysuccinimide (NHS) solution and a 390 mM 1-ethyl-3-(3-dimethylaminopropyl)-carbodiimide (EDC) hydrochloride salt solution in water. Both proteins were diluted to0.05 mg/mL in 20 mM HEPES pH 6.0 for immobilization onto different flow cells of the chip. After a 5 min injection of the protein, remaining reactive groups were deactivated by injecting a 1 M ethanolamine hydrochloride solution in aqueous NaOH (pH 8.5) for 5 min. Different chips were used with immobilization levels ranging between 3,000 and 5,000 response units (RU). To determine kinetic parameters for the binding of compound 6 to human FD and FB, several independent experiments were run. Threefold serial dilutions of compound 6 were prepared ranging from 0.4 to 900 nM. Each solution was injected for 60 s at a flow rate of 30 to 60 μL/min with a dissociation time of at least 1,200 s (parameters identical within one experiment).
- SPR Binding Affinity Assay (Standard) Standard kinetics: an independent association-dissociation cycle was run for each concentration by injecting compound solution and waiting for dissociation before the injection of the following concentration. SPR experiments were performed at 25 °C using a Biacore T200 instrument (GE Healthcare). PBS (pH 7.4) supplemented with 0.05% Tween 20 was used as running buffer. Human FD and FB were immobilized covalently to a Series S Sensor Chip CM5 (GE Healthcare) at a flow rate of 10 μL/min using an amine-coupling protocol. Reagents for the immobilization were purchased from GE Healthcare (Amine Coupling Kit; BR-1000-50). The sensor-chip surface was activated by a 5 min injection of a 1:1 (v/v) mixture of a 100 mM N-hydroxysuccinimide (NHS) solution and a 390 mM 1-ethyl-3-(3-dimethylaminopropyl)-carbodiimide (EDC) hydrochloride salt solution in water. Both proteins were diluted to0.05 mg/mL in 20 mM HEPES pH 6.0 for immobilization onto different flow cells of the chip. After a 5 min injection of the protein, remaining reactive groups were deactivated by injecting a 1 M ethanolamine hydrochloride solution in aqueous NaOH (pH 8.5) for 5 min. Different chips were used with immobilization levels ranging between 3,000 and 5,000 response units (RU). To determine kinetic parameters for the binding of compound 6 to human FD and FB, several independent experiments were run. Threefold serial dilutions of compound 6 were prepared ranging from 0.4 to 900 nM. Each solution was injected for 60 s at a flow rate of 30 to 60 μL/min with a dissociation time of at least 1,200 s (parameters identical within one experiment).
- SPR Assay Surface plasmon resonance data was collected on a Biacore T200 or 3000 system (GE Healthcare) at 25� C. Streptavidin was immobilized on a CM5 (GE Healthcare) or CMD500d sensor chip (XanTec Bioanalytics) using standard amine-coupling chemistry at 25� C. with HBS-N (10 mM HEPES, 0.15 M NaCl, pH 7.4) as the running buffer. Briefly, the carboxymethyl dextran surface was activated with a 12 min injection of a 1:1 ratio of 0.4 M 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (EDC)/0.1 M N-hydroxy succinimide (NHS) at a flow rate of 10 μL/min. For capture of streptavidin, protein was diluted to 0.2 mg/mL in 10 mM sodium acetate (pH 4.5) and captured by injecting 100 μL onto the activated chip surface. Residual activated groups were blocked with a 7 min injection of 1 M ethanolamine (pH 8.5). Avi-tagged PCSK9 protein was captured on the streptavidin surface by injection of 150 μL of protein diluted to 16 pg/mL in FIBS-N, 0.05% tween-20, 0.1 mM CaCl2. Typical surface densities obtained were 8000-10000 RU. SPR binding data were obtained using an appropriate dilution series of each compound at a flow rate of 30 μL/min, with a capture time of 100 s and dissociation times of 300 s. Running buffer for compound binding studies was FIBS-N, 0.05% tween-20, 0.1 mM CaCl2), 4% DMSO. Data were corrected for DMSO excluded volume effects. All data were double-referenced for blank injections and reference surface using standard processing procedures and data processing and kinetic fitting were performed using Scrubber software, version 2.0c (BioLogic Software).
- Surface Plasmon Resonance (SPR) Assay Surface plasmon resonance data was collected on a Biacore T200 or 3000 system (GE Healthcare) at 25� C. Streptavidin was immobilized on a CMS (GE Healthcare) or CMD500d sensor chip (XanTec Bioanalytics) using standard amine-coupling chemistry at 25� C. with HBS-N (10 mM HEPES, 0.15 M NaCl, pH 7.4) as the running buffer. Briefly, the carboxymethyl dextran surface was activated with a 12 min injection of a 1:1 ratio of 0.4 M 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (EDC)/0.1 M N-hydroxy succinimide (NHS) at a flow rate of 10 μL/min. For capture of streptavidin, protein was diluted to 0.2 mg/mL in 10 mM sodium acetate (pH 4.5) and captured by injecting 100 μL onto the activated chip surface. Residual activated groups were blocked with a 7 min injection of 1 M ethanolamine (pH 8.5). Avi-tagged PCSK9 protein was captured on the streptavidin surface by injection of 150 μL of protein diluted to 16 μg/mL in HBS-N, 0.05% tween-20, 0.1 mM CaCl2. Typical surface densities obtained were 8000-10000 RU. SPR binding data were obtained using an appropriate dilution series of each compound at a flow rate of 30 μL/ min, with a capture time of 100 s and dissociation times of 300 s. Running buffer for compound binding studies was HBS-N, 0.05% tween-20, 0.1 mM CaCl2, 4% DMSO. Data were corrected for DMSO excluded volume effects. All data were double-referenced for blank injections and reference surface using standard processing procedures and data processing and kinetic fitting were performed using Scrubber software, version 2.0c (BioLogic Software).
- [3H]-AEA Cellular Uptake Screening for AEA cellular uptake inhibition was performed in a semi-automated procedure: Pipetting and washing steps were performed by a Biomek3000 laboratory workstation. First, required amounts of U937 cells were centrifuged at 100�g for 5 min and resuspended in RPMI (37� C.) to a final concentration of 2�106 cells/mL. Then, 250 μL of cell suspension (0.5�106 cells per sample) were transferred into AquaSil silanized glass vials (Chromacol 1.1-MTV) in 96-well format. After addition of 5 μL vehicle (DMSO) or compounds the cells were incubated at 37� C. for 15 min. As positive controls OMDM-2 and UCM707 were used at 10 μM in each run. The ETI-T compounds were measured at up to 7 concentrations in triplicates from 100 pM-100 μM. After pre-incubation, a mixture of 0.5 nM[ethanolamine-1-3H]-AEA, (60 Ci/mmol) and 99.5 nM of cold AEA (final 100 nM) was added and samples were incubated at 37� C. for another 15 min. The reaction was stopped by rapid filtration over UniFilter-96 GF/C filters (PerkinElmer) pre-soaked with PBS 0.25% BSA. Cells were washed three times with 100 μL ice-cold PBS buffer containing 1% fatty acid free BSA. After drying, 45 μL MicroScint 20 scintillation cocktail (PerkinElmer, Waltham, Mass., US) was added to the wells and the plate was sealed. Radioactivity was measured by liquid scintillation counting on a PerkinElmer Wallac Trilux MicroBeta 1450 during 2 min. Non-specific binding of [3H]AEA (100 nM) to the glass vials was never higher than 10%. IC50 values were calculated by GraphPad by non-linear regression using the built-in log(inhibitor) vs. response-variable slope (four parameters) function.
- In Vitro Radioactive Assays of CETP-Catalyzed CE and TG Transfer (RTA Assay) Radiolabeled donor particles are generated by first combining 100 μl of 200 μM butylated hydroxyl toluene in CHCl3, 216 μL of 21.57 mM DOPC in EtOH, and either 500 μCi [3H]-triolein (Perkin Elmer #NET-431) or 500 μCi [3H]-cholesteryl oleate (GE #TRK886) in a glass tube. Reagents are mixed, dried under nitrogen, and then resuspended in 2 mL of 50 mM Tris, 27 μM EDTA at pH 7.4. After a brief vortex, the solution is sonicated until clear and mixed with 20 mL of fresh human serum. The mixture is incubated overnight at 37° C. The [3H] labeled LDL substrate is separated at 1.063 g/ml density by sequential ultracentrifugal flotation in NaBr according to the method by (Havel, Eder et al. 1955; Chapman, Goldstein et al. 1981). Once isolated the particles are dialyzed 3× in CETP buffer (50 mM Tris, pH 7.4, 100 mM NaCl, 1 mM EDTA). Human HDL is purchased from Intracel and used as the acceptor particles. Transfer assays are performed in a 96 or 384-well v-bottom polypropylene plate. For the RTA using recombinant CETP (2% RTA), an assay cocktail is prepared with the final concentrations 128 μg/mL HDL, 20 nM rCETP, 2% human serum, and 1×CETP buffer. 1 μL of each test compound diluted in DMSO is added to 47 μL of assay cocktail per well and incubated at 37° C. for 1 hour. To initiate the transfer reaction, 2 μL radiolabeled LDL is added. After an additional 60 min of incubation at 37° C., the transfer action is terminated by precipitation of LDL with an equal volume of 20% W/V PEG 8000. The plates are centrifuged at 2000 rpm for 30 minutes at 4° C. A 40 μL aliquot of the HDL-containing supernatant is transferred to a Packard Optiplate™ with 200 μL of MicroScint™ 20. After mixing, plates are counted by liquid scintillation.
- Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org Lett 2006, 8 (10), 2023) was labeled with MR121 fluorophore (CAS 185308-24-1, 1-(3-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2′,3′-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid.Assay working solutions were made as follows:Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaCl2, 1 mM MgCl2, 0.01% Triton-X-100, pH 8.0; ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5� final concentration in assay buffer; MR121 substrate solution: MR121 substrate stock solution (800 μM MR121 substrate in DMSO), diluted to 2-5� final concentration in assay buffer.Test compounds (10 mM stock in DMSO, 8 μL) were obtained in 384 well sample plates (Corning Costar #3655) and diluted with 8 μL DMSO. Row-wise serial dilutions were made by transferring 8 μL cpd solution to the next row up to row O. The compound and control solutions were mixed five times and 2 L were transferred to 384 well assay plates (Corning Costar #3702). Then, 15 μL of 41.7 nM ATX solution was added (30 nM final concentration), mixed five times and then incubated for 15 minutes at 30� C. 10 μL of MR121 substrate solution was added (1 μM final concentration), mixed 30 times and then incubated for 15 minutes at 30� C. Fluorescence was then measured every 2 minutes for 1 hour (Perkin Elmer plate: vision multimode reader); light intensity: 2.5%; exp. time: 1.4 sec, Filter: Fluo_630/690 nm).
- Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org Lett 2006, 8 (10), 2023) was labeled with MR121 fluorophore (CAS 185308-24-1, 143-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2',3'-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid.Assay working solutions were made as follows:Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaCl2, 1 mM MgCl2, 0.01% Triton-X-100, pH 8.0;ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5x final concentration in assay buffer. MR121 substrate solution: MR121 substrate stock solution (800 uM MR121 substrate in DMSO), diluted to 2-5x final concentration in assay buffer. Test compounds (10 mM stock in DMSO, 8 uL) were obtained in 384 well sample plates (Corning Costar #3655) and diluted with 8 uL DMSO. Row-wise serial dilutions were made by transferring 8 uL cpd solution to the next row up to row 0. The compound and control solutions were mixed five times and 2 uL were transferred to 384 well assay plates (Corning Costar #3702). Then, 15 uL of 41.7 nM ATX solution was added (30 nM final concentration), mixed five times and then incubated for 15 minutes at 30° C. 10 uL of MR121 substrate solution was added (1 uM final concentration), mixed 30 times and then incubated for 15 minutes at 30° C. Fluorescence was then measured every 2 minutes for 1 hour (Perkin Elmer plate: vision multimode reader); light intensity: 2.5%; exp. time: 1.4 sec, Filter: Fluo_630/690 nm) and IC50 values were calculated from these readouts.
- Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org Lett 2006, 8 (10), 2023) was labeled with MR121 fluorophore (CAS 185308-24-1,1-(3-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2′,3′-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid.Assay working solutions were made as follows:Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaCl2, 1 mM MgCl2, 0.01%Triton-X-100, pH 8.0;ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5�final concentration in assay buffer;MR121 substrate solution: MR121 substrate stock solution (800 M MR121 substrate in DMSO), diluted to 2-5�final concentration in assay buffer.Test compounds (10 mM stock in DMSO, 8 μL) were obtained in 384 well sample plates (Corning Costar #3655) and diluted with 8 μL DMSO. Row-wise serial dilutions were made by transferring 8 μL cpd solution to the next row up to row O. The compound and control solutions were mixed five times and 2 μL were transferred to 384 well assay plates (Corning Costar #3702). Then, 15 μL of 41.7 nM ATX solution was added (30 nM final concentration), mixed five times and then incubated for 15 minutes at 30� C. 10 μL of MR121 substrate solution was added (1 μM final concentration), mixed 30 times and then incubated for 15 minutes at 30� C. Fluorescence was then measured every 2 minutes for 1 hour (Perkin Elmer plate: vision multimode reader); light intensity: 2.5%; exp. time: 1.4 sec, Filter: Fluo_630/690 nm) and IC50 values were calculated from these readouts.
- Human ATX Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org Lett 2006, 8 (10), 2023) was labeled with MR121 fluorophore (CAS 185308-24-1, 1-(3-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2′,3′-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid. Assay working solutions were made as follows: Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaC2, 1 mM MgCl2, 0.01% Triton-X-100, pH 8.0; ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5� final concentration in assay buffer; MR121 substrate solution: MR121 substrate stock solution (800 μM MR121 substrate in DMSO), diluted to 2-5� final concentration in assay buffer. Test compounds (10 mM stock in DMSO, 8 μL) were obtained in 384 well sample plates (Corning Costar #3655) and diluted with 8 μL DMSO. Row-wise serial dilutions were made by transferring 8 μL cpd solution to the next row up to row O. The compound and control solutions were mixed five times and 2 μL were transferred to 384 well assay plates (Corning Costar #3702). Then, 15 μL of 41.7 nM ATX solution was added (30 nM final concentration), mixed five times and then incubated for 15 minutes at 30� C. 10 μL of MR121 substrate solution was added (1 μM final concentration), mixed 30 times and then incubated for 15 minutes at 30� C. Fluorescence was then measured every 2 minutes for 1 hour (Perkin Elmer plate: vision multimode reader); light intensity: 2.5%; exp. time: 1.4 sec, Filter: Fluo_630/690 nm) and IC50 values were calculated from these readouts.
- Human ATX Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org Lett 2006, 8 (10), 2023) was labeled with MR121 fluorophore (CAS 185308-24-1, 1-(3-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2′,3′-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid.Assay working solutions were made as follows:Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaCl2, 1 mM MgCl2, 0.01% Triton-X-100, pH 8.0;ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5� final concentration in assay buffer;MR121 substrate solution: MR121 substrate stock solution (800 μM MR121 substrate in DMSO), diluted to 2-5� final concentration in assay buffer.Test compounds (10 mM stock in DMSO, 8 μL) were obtained in 384 well sample plates (Corning Costar #3655) and diluted with 8 μL DMSO. Row-wise serial dilutions were made by transferring 8 μL cpd solution to the next row up to row 0. The compound and control solutions were mixed five times and 2 μL were transferred to 384 well assay plates (Corning Costar #3702). Then, 15 μL of 41.7 nM ATX solution was added (30 nM final concentration), mixed five times and then incubated for 15 minutes at 30� C. 10 μL of MR121 substrate solution was added (1 μM final concentration), mixed 30 times and then incubated for 15 minutes at 30� C. Fluorescence was then measured every 2 minutes for 1 hour (Perkin Elmer plate: vision multimode reader); light intensity: 2.5%; exp. time: 1.4 sec, Filter: Fluo_630/690 nm) and IC50 values were calculated from these readouts.
- Human ATX Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org Lett 2006, 8 (10), 2023) was labeled with MR121 fluorophore (CAS 185308-24-1, 1-(3-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2′,3′-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid.Assay working solutions were made as follows:Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaCl2, 1 mM MgCl2, 0.01% Triton-X-100, pH 8.0;ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5� final concentration in assay buffer;MR121 substrate solution: MR121 substrate stock solution (800 μM MR121 substrate in DMSO), diluted to 2-5� final concentration in assay buffer.Test compounds (10 mM stock in DMSO, 8 μL) were obtained in 384 well sample plates (Corning Costar #3655) and diluted with 8 μL DMSO. Row-wise serial dilutions were made by transferring 8 μL cpd solution to the next row up to row O. The compound and control solutions were mixed five times and 2 μL were transferred to 384 well assay plates (Corning Costar #3702). Then, 15 μL of 41.7 nM ATX solution was added (30 nM final concentration), mixed five times and then incubated for 15 minutes at 30� C. 10 μL of MR121 substrate solution was added (1 μM final concentration), mixed 30 times and then incubated for 15 minutes at 30� C. Fluorescence was then measured every 2 minutes for 1 hour (Perkin Elmer plate: vision multimode reader); light intensity: 2.5%; exp. time: 1.4 sec, Filter: Fluo_630/690 nm) and IC50 values were calculated from these readouts.
- Human ATX Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org. Lett. 2006, 8, 2023) was labeled with MR121 fluorophore [CAS RN 185308-24-1], 1-(3-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2′,3′-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid.Assay working solutions were made as follows:Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaCl2, 1 mM MgCl2, 0.01% Triton-X-100, pH 8.0;ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5� final concentration in assay buffer;MR121 substrate solution: MR121 substrate stock solution (800 μM MR121 substrate in DMSO), diluted to 2-5� final concentration in assay buffer.Test compounds (10 mM stock in DMSO, 8 μL) were obtained in 384 well sample plates (Corning Costar #3655) and diluted with 8 μL DMSO. Row-wise serial dilutions were made by transferring 8 μL cpd solution to the next row up to row O. The compound and control solutions were mixed five times and 2 μL were transferred to 384 well assay plates (Corning Costar #3702). Then, 15 μL of 41.7 nM ATX solution was added (30 nM final concentration), mixed five times and then incubated for 15 minutes at 30� C. 10 μL of MR121 substrate solution was added (1 μM final concentration), mixed 30 times and then incubated for 15 minutes at 30� C. Fluorescence was then measured every 2 min for 1 h (Perkin Elmer plate: vision multimode reader); light intensity: 2.5%; exp. time: 1.4 s, Filter: Fluo_630/690 nm) and IC50 values were calculated from these readouts.
- Human Atx Enzyme Inhibition Assay ATX inhibition was measured by a fluorescence quenching assay using a specifically labeled substrate analogue (MR121 substrate). To obtain this MR121 substrate, BOC and TBS protected 6-amino-hexanoic acid (R)-3-({2-[3-(2-{2-[2-(2-amino-ethoxy)-ethoxy]-ethoxy}-ethoxy)-propionylamino]-ethoxy}-hydroxy-phosphoryloxy)-2-hydroxy-propyl ester (Ferguson et al., Org Lett 2006, 8 (10), 2023) was labeled with MR121 fluorophore (CAS 185308-24-1,1-(3-carboxypropyl)-11-ethyl-1,2,3,4,8,9,10,11-octahydro-dipyrido[3,2-b:2′,3′-i]phenoxazin-13-ium) on the free amine of the ethanolamine side and then, after deprotection, subsequently with tryptophan on the side of the aminohexanoic acid.Assay working solutions were made as follows:Assay buffer (50 mM Tris-HCl, 140 mM NaCl, 5 mM KCl, 1 mM CaCl2, 1 mM MgCl2, 0.01% Triton-X-100, pH 8.0;ATX solution: ATX (human His-tagged) stock solution (1.08 mg/mL in 20 mM bicine, pH 8.5, 0.15 M NaCl, 10% glycerol, 0.3% CHAPS, 0.02% NaN3), diluted to 1.4-2.5� final concentration in assay buffer;MR121 substrate solution: MR121 substrate stock solution (800 μM MR121 substrate in DMSO), diluted to 2-5� final concentration in assay buffer.Test compounds (10 mM stock in DMSO, 8 μL) were obtained in 384 well sample plates (Corning Costar #3655) and diluted with 8 μL DMSO. Row-wise serial dilutions were made by transferring 8 μL cpd solution to the next row up to row O. The compound and control solutions were mixed five times and 2 μL were transferred to 384 well assay plates (Corning Costar #3702). Then, 15 μL of 41.7 nM ATX solution was added (30 nM final concentration), mixed five times and then incubated for 15 minutes at 30� C. 10 μL of MR121 substrate solution was added (1 M final concentration), mixed 30 times and then incubated for 15 minutes at 30� C. Fluorescence was then measured every 2 minutes for 1 hour (Perkin Elmer plate: vision multimode reader); light intensity: 2.5%; exp. time: 1.4 sec, Filter: Fluo_630/690 nm) and IC50 values were calculated from these readouts.
- Determining Endocannabinoid Hydrolase Activity In the Experimental example, endocannabinoid hydrolases used were Fatty Acid Amide Hydrolase (FAAH) and N-acylethanolamide hydrolyzing acid amidase (NAAA), which were prepared by the method described in the document (PMCID: PMC3423427, PMC3723234, PMC2692831, PMC3382457). The preparation method was as followed: a plasmid (pCDNA3.1/NAAA or pCDNA3.1/FAAH) carrying a whole NAAA/FAAH gene was constructed, wherein the plasmid carried Cytomegalovirus (CMV) promoter and Neomycin selectable gene; the plasmid was transformed into HEK-293 cell via lipid medium, stable cell lines expressing NAAA/FAAH at a high level were obtained by G418 screening and Western-blot method. HEK-293 recombinant cells were cultured and collected, washed with PBS for 2-3 times, and ultrasonically treated in 20 mM Tris-HCl containing 0.32 M sucrose, then repeatedly frozen and thawed twice, and then centrifuged at 4� C., 800 g for 15 min. The supernatant (i.e., the desired protein) was collected, the protein concentration was determined by BCA method, and the protein was diluted to a concentration of 1 mg/mL, and sub-packaged and stored in a refrigerator at −80� C. for further use.In the Experimental example, PBS solution used was prepared as followed: 8 g NaCl, 0.2 g KCl, 1.44 g Na2HPO4, and 0.24 g KH2PO4 were dissolved in 1 L ultrapure water, and the resultant solution was subjected to moist heat sterilization and stored at 4� C.30 μL (1 mg/mL) endocannabinoid hydrolase was added to a sample vial, and 2 μL DMSO (Blank control group) or a different concentration of a test compound (Compounds 1-46 prepared in the Examples according to the invention) was further added. The reaction was carried out at 37� C. for 10 min. 170 μL buffer (the buffer consisted of 50 mM disodium Hydrogen Phosphate, 0.1% Triton X-100, 3 mM DTT, 150 μL) containing enzymatic hydrolysis substrate (the substrate was a heptadecenoyl ethanolamine containing a double bond and 17 carbon atoms, abbreviated as 17:1 FAE) was further added, wherein the concentration of 17:1 FAE was 5 μM. The reaction was carried out at 37� C. for 30 min, and 200 μL methanol solution containing internal standard (the internal standard was margaric acid, at a concentration of 1 nmol) was then added to stop the reaction. LC-MS was used to determine the yield of the hydrolysate 17:1 FA (i.e., a heptadecenoic acid containing a double bond) of 17:1 FAE, then a graph was plotted with Graphpad Prism 5. Thereby, the IC50 of the test compound on endocannabinoid hydrolase was determined.By the method above, the inhibitory effects of the Compounds 1-46 prepared in the invention on NAAA and FAAH were determined. The results were shown in Table 1, wherein IC50 (NAAA) represents a concentration that inhibits NAAA activity to 50% of the activity prior to inhibition, IC50 (FAAH) represents a concentration that inhibits FAAH activity to 50% of the activity prior to inhibition, and >100 μM represents that IC50 of a compound on a corresponding enzyme is above 100 μM, indicating that the compound has no inhibitory effect on the enzyme.
 BDBM246504 4-Methylumbelliferyl oleate (4-MU oleate)
BDBM246504 4-Methylumbelliferyl oleate (4-MU oleate) BDBM237193 Rutin oleate
BDBM237193 Rutin oleate BDBM50448437 CHEMBL3122151 Oleate
BDBM50448437 CHEMBL3122151 Oleate ethanolamine BDBM7973 CHEMBL104943 2-aminoethan-1-ol
ethanolamine BDBM7973 CHEMBL104943 2-aminoethan-1-ol 2-(benzylamino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 1 BDBM18072
2-(benzylamino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 1 BDBM18072 1-(3-bromophenyl)-2-{[(2-methoxyphenyl)methyl]amino}ethan-1-ol BDBM18082 ethanolamine, 18
1-(3-bromophenyl)-2-{[(2-methoxyphenyl)methyl]amino}ethan-1-ol BDBM18082 ethanolamine, 18 2-{[(2-methoxyphenyl)methyl]amino}-1-(3-methylphenyl)ethan-1-ol BDBM18083 ethanolamine, 19
2-{[(2-methoxyphenyl)methyl]amino}-1-(3-methylphenyl)ethan-1-ol BDBM18083 ethanolamine, 19 BDBM18077 ethanolamine, 9 2-[({2-hydroxy-2-[3-(trifluoromethyl)phenyl]ethyl}amino)methyl]phenol
BDBM18077 ethanolamine, 9 2-[({2-hydroxy-2-[3-(trifluoromethyl)phenyl]ethyl}amino)methyl]phenol 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-chlorophenyl)ethan-1-ol ethanolamine, 22 BDBM18085
2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-chlorophenyl)ethan-1-ol ethanolamine, 22 BDBM18085 2-[({2-hydroxy-2-[3-(trifluoromethyl)phenyl]ethyl}amino)methyl]benzoic acid BDBM18075 ethanolamine, 7
2-[({2-hydroxy-2-[3-(trifluoromethyl)phenyl]ethyl}amino)methyl]benzoic acid BDBM18075 ethanolamine, 7 BDBM18074 ethanolamine, 6 2-{[(2-methylphenyl)methyl]amino}-1-[3-(trifluoromethyl)phenyl]ethan-1-ol
BDBM18074 ethanolamine, 6 2-{[(2-methylphenyl)methyl]amino}-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 11 2-{[(2-methoxyphenyl)methyl]amino}-1-[3-(trifluoromethyl)phenyl]ethan-1-ol BDBM18079
ethanolamine, 11 2-{[(2-methoxyphenyl)methyl]amino}-1-[3-(trifluoromethyl)phenyl]ethan-1-ol BDBM18079 ethanolamine, 20 BDBM18084 2-{[(2-methoxyphenyl)methyl]amino}-1-[3-(trifluoromethoxy)phenyl]ethan-1-ol
ethanolamine, 20 BDBM18084 2-{[(2-methoxyphenyl)methyl]amino}-1-[3-(trifluoromethoxy)phenyl]ethan-1-ol ethanolamine, 23 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-iodophenyl)ethan-1-ol BDBM18086
ethanolamine, 23 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-iodophenyl)ethan-1-ol BDBM18086 ethanolamine, 24 BDBM18087 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-methoxyphenyl)ethan-1-ol
ethanolamine, 24 BDBM18087 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(3-methoxyphenyl)ethan-1-ol oleicacid (Z)-9-octadecenoic acid OLEATE BDBM50150484 Octadec-9-enoic acid (Z)-Octadec-9-enoic acid CHEMBL8659 oleaic acid
oleicacid (Z)-9-octadecenoic acid OLEATE BDBM50150484 Octadec-9-enoic acid (Z)-Octadec-9-enoic acid CHEMBL8659 oleaic acid 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-[3-(hydroxymethyl)phenyl]ethan-1-ol BDBM18088 ethanolamine, 25
2-({[2-(benzyloxy)phenyl]methyl}amino)-1-[3-(hydroxymethyl)phenyl]ethan-1-ol BDBM18088 ethanolamine, 25 2-({[2-(difluoromethoxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 10 BDBM18078
2-({[2-(difluoromethoxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 10 BDBM18078 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]acetonitrile BDBM18092 ethanolamine, 37
2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]acetonitrile BDBM18092 ethanolamine, 37 BDBM18104 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]acetamide ethanolamine, 49
BDBM18104 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]acetamide ethanolamine, 49 ethanolamine, 13 BDBM18081 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol
ethanolamine, 13 BDBM18081 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol ethanolamine, 34 1-(4-bromothiophen-2-yl)-2-{[(2-methoxyphenyl)methyl]amino}ethan-1-ol BDBM18089
ethanolamine, 34 1-(4-bromothiophen-2-yl)-2-{[(2-methoxyphenyl)methyl]amino}ethan-1-ol BDBM18089 ethanolamine, 35 BDBM18090 1-(4-bromothiophen-2-yl)-2-{[(2-phenoxyphenyl)methyl]amino}ethan-1-ol
ethanolamine, 35 BDBM18090 1-(4-bromothiophen-2-yl)-2-{[(2-phenoxyphenyl)methyl]amino}ethan-1-ol 3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]propanoic acid BDBM18108 ethanolamine, 53
3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]propanoic acid BDBM18108 ethanolamine, 53 BDBM18091 1-(4-bromothiophen-2-yl)-2-({[2-(methoxymethoxy)phenyl]methyl}amino)ethan-1-ol ethanolamine, 36
BDBM18091 1-(4-bromothiophen-2-yl)-2-({[2-(methoxymethoxy)phenyl]methyl}amino)ethan-1-ol ethanolamine, 36 BDBM18105 ethanolamine, 50 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-ethylacetamide
BDBM18105 ethanolamine, 50 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-ethylacetamide ethanolamine, 43 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol BDBM18098
ethanolamine, 43 2-({[2-(benzyloxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol BDBM18098 ethanolamine, 56 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]butanoic acid BDBM18111
ethanolamine, 56 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]butanoic acid BDBM18111 2-({[2-(morpholin-4-yl)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol BDBM18076 ethanolamine, 8
2-({[2-(morpholin-4-yl)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol BDBM18076 ethanolamine, 8 BDBM18093 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol ethanolamine, 38
BDBM18093 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol ethanolamine, 38 BDBM18094 ethanolamine, 39 2-({[2-(2-aminoethoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol
BDBM18094 ethanolamine, 39 2-({[2-(2-aminoethoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol BDBM18114 2-({[2-(4-aminobutoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol ethanolamine, 59
BDBM18114 2-({[2-(4-aminobutoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol ethanolamine, 59 1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-2-ylmethoxy)phenyl]methyl}amino)ethan-1-ol ethanolamine, 45 BDBM18100
1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-2-ylmethoxy)phenyl]methyl}amino)ethan-1-ol ethanolamine, 45 BDBM18100 1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-4-ylmethoxy)phenyl]methyl}amino)ethan-1-ol BDBM18102 ethanolamine, 47
1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-4-ylmethoxy)phenyl]methyl}amino)ethan-1-ol BDBM18102 ethanolamine, 47 BDBM18106 ethanolamine, 51 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]acetamide
BDBM18106 ethanolamine, 51 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]acetamide BDBM18116 ethanolamine, 64 2-[2-({[(2R)-2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol
BDBM18116 ethanolamine, 64 2-[2-({[(2R)-2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol BDBM18117 ethanolamine, 65 2-[2-({[(2S)-2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol
BDBM18117 ethanolamine, 65 2-[2-({[(2S)-2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethan-1-ol ethanolamine, 44 BDBM18099 1-(4-bromothiophen-2-yl)-2-[({2-[(4-methanesulfonylphenyl)methoxy]phenyl}methyl)amino]ethan-1-ol
ethanolamine, 44 BDBM18099 1-(4-bromothiophen-2-yl)-2-[({2-[(4-methanesulfonylphenyl)methoxy]phenyl}methyl)amino]ethan-1-ol ethanolamine, 46 1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-3-ylmethoxy)phenyl]methyl}amino)ethan-1-ol BDBM18101
ethanolamine, 46 1-(4-bromothiophen-2-yl)-2-({[2-(pyridin-3-ylmethoxy)phenyl]methyl}amino)ethan-1-ol BDBM18101 ethanolamine, 54 3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]propanamide BDBM18109
ethanolamine, 54 3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]propanamide BDBM18109 ethanolamine, 57 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]butanamide BDBM18112
ethanolamine, 57 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-[(methylcarbamoyl)methyl]butanamide BDBM18112 3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylpropanamide ethanolamine, 55 BDBM18110
3-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylpropanamide ethanolamine, 55 BDBM18110 4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylbutanamide BDBM18113 ethanolamine, 58
4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylbutanamide BDBM18113 ethanolamine, 58 BDBM18107 ethanolamine, 52 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylacetamide
BDBM18107 ethanolamine, 52 2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]-N-(2-hydroxyethyl)-N-methylacetamide ethanolamine, 12 BDBM18080 2-({[2-(prop-2-en-1-yloxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol
ethanolamine, 12 BDBM18080 2-({[2-(prop-2-en-1-yloxy)phenyl]methyl}amino)-1-[3-(trifluoromethyl)phenyl]ethan-1-ol BDBM18103 ethanolamine, 48 2-({[2-(1H-1,3-benzodiazol-2-ylmethoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol
BDBM18103 ethanolamine, 48 2-({[2-(1H-1,3-benzodiazol-2-ylmethoxy)phenyl]methyl}amino)-1-(4-bromothiophen-2-yl)ethan-1-ol ethanolamine, 41 1-(4-bromothiophen-2-yl)-2-[({2-[2-(morpholin-4-yl)ethoxy]phenyl}methyl)amino]ethan-1-ol BDBM18096
ethanolamine, 41 1-(4-bromothiophen-2-yl)-2-[({2-[2-(morpholin-4-yl)ethoxy]phenyl}methyl)amino]ethan-1-ol BDBM18096 ethanolamine, 42 BDBM18097 1-(4-bromothiophen-2-yl)-2-({[2-(prop-2-en-1-yloxy)phenyl]methyl}amino)ethan-1-ol
ethanolamine, 42 BDBM18097 1-(4-bromothiophen-2-yl)-2-({[2-(prop-2-en-1-yloxy)phenyl]methyl}amino)ethan-1-ol 2-amino-ethanol dihydrogen phosphate O-phosphocolamine ethanolamine O-phosphate OPE phosphonoethanolamine monoaminoethyl phosphate colaminphosphoric acid 2-amino-ethanol phosphate PEA colamine phosphate mono(2-aminoethyl) phosphate colamine phosphoric acid 2-aminoethyl dihydrogen phosphate phosphoric acid 2-aminoethyl phenyl ester CHEMBL146972 BDBM50281572 EAP ethanolamine acid phosphate pEtN O-phosphoethanolamine
2-amino-ethanol dihydrogen phosphate O-phosphocolamine ethanolamine O-phosphate OPE phosphonoethanolamine monoaminoethyl phosphate colaminphosphoric acid 2-amino-ethanol phosphate PEA colamine phosphate mono(2-aminoethyl) phosphate colamine phosphoric acid 2-aminoethyl dihydrogen phosphate phosphoric acid 2-aminoethyl phenyl ester CHEMBL146972 BDBM50281572 EAP ethanolamine acid phosphate pEtN O-phosphoethanolamine BDBM18115 ethanolamine, 60 2-{4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]butyl}-2,3-dihydro-1H-isoindole-1,3-dione
BDBM18115 ethanolamine, 60 2-{4-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]butyl}-2,3-dihydro-1H-isoindole-1,3-dione ethanolamine, 40 BDBM18095 2-{2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethyl}-2,3-dihydro-1H-isoindole-1,3-dione
ethanolamine, 40 BDBM18095 2-{2-[2-({[2-(4-bromothiophen-2-yl)-2-hydroxyethyl]amino}methyl)phenoxy]ethyl}-2,3-dihydro-1H-isoindole-1,3-dione MLS002153867 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methyl-2-pyridinone BDBM66087 cid_38911 2-azanylethanol;6-cyclohexyl-4-methyl-1-oxidanyl-pyridin-2-one Ciclopirox Ciclopirox ethanolamine SMR001233223 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methyl-2-pyridone 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methylpyridin-2-one
MLS002153867 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methyl-2-pyridinone BDBM66087 cid_38911 2-azanylethanol;6-cyclohexyl-4-methyl-1-oxidanyl-pyridin-2-one Ciclopirox Ciclopirox ethanolamine SMR001233223 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methyl-2-pyridone 2-aminoethanol;6-cyclohexyl-1-hydroxy-4-methylpyridin-2-one